You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000222_03394
You are here: Home > Sequence: MGYG000000222_03394
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides sp003473295 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp003473295 | |||||||||||
| CAZyme ID | MGYG000000222_03394 | |||||||||||
| CAZy Family | CE7 | |||||||||||
| CAZyme Description | Acetyl esterase Axe7A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14987; End: 16264 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE7 | 127 | 420 | 7.8e-90 | 0.9648562300319489 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3458 | Axe1 | 2.17e-52 | 119 | 421 | 1 | 315 | Cephalosporin-C deacetylase or related acetyl esterase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam05448 | AXE1 | 1.88e-50 | 136 | 421 | 20 | 315 | Acetyl xylan esterase (AXE1). This family consists of several bacterial acetyl xylan esterase proteins. Acetyl xylan esterases are enzymes that hydrolyze the ester linkages of the acetyl groups in position 2 and/or 3 of the xylose moieties of natural acetylated xylan from hardwood. These enzymes are one of the accessory enzymes which are part of the xylanolytic system, together with xylanases, beta-xylosidases, alpha-arabinofuranosidases and methylglucuronidases; these are all required for the complete hydrolysis of xylan. |
| COG1506 | DAP2 | 2.40e-06 | 183 | 417 | 375 | 605 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| COG0412 | DLH | 2.74e-05 | 182 | 322 | 10 | 146 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT49362.1 | 3.09e-251 | 1 | 420 | 1 | 420 |
| QRM99790.1 | 1.20e-232 | 1 | 420 | 1 | 420 |
| QUT31430.1 | 1.20e-232 | 1 | 420 | 1 | 420 |
| BCG54171.1 | 1.39e-213 | 1 | 422 | 3 | 425 |
| QUT54228.1 | 9.16e-171 | 1 | 421 | 1 | 423 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6AGQ_A | 3.60e-38 | 119 | 406 | 1 | 303 | Acetylxylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4],6AGQ_B Acetyl xylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4],6AGQ_C Acetyl xylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4],6AGQ_D Acetyl xylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4],6AGQ_E Acetyl xylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4],6AGQ_F Acetyl xylan esterase from Paenibacillus sp. R4 [Paenibacillus sp. R4] |
| 1ODS_A | 1.10e-29 | 130 | 425 | 15 | 317 | CephalosporinC deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_B Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_C Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_D Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_E Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_F Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_G Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis],1ODS_H Cephalosporin C deacetylase from Bacillus subtilis [Bacillus subtilis] |
| 1ODT_C | 2.90e-29 | 130 | 425 | 15 | 317 | cephalosporinC deacetylase mutated, in complex with acetate [Bacillus subtilis],1ODT_H cephalosporin C deacetylase mutated, in complex with acetate [Bacillus subtilis] |
| 1L7A_A | 2.90e-29 | 130 | 422 | 15 | 314 | structuralGenomics, crystal structure of Cephalosporin C deacetylase [Bacillus subtilis],1L7A_B structural Genomics, crystal structure of Cephalosporin C deacetylase [Bacillus subtilis] |
| 5HFN_A | 4.82e-29 | 130 | 406 | 27 | 289 | Crystalstructure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8],5HFN_B Crystal structure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8],5HFN_C Crystal structure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8],5HFN_D Crystal structure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8],5HFN_E Crystal structure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8],5HFN_F Crystal structure of a loop truncation variant of Thermotoga maritima Acetyl Esterase TM0077 (apo structure) at 2.75 Angstrom resolution [Thermotoga maritima MSB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EXI2 | 1.14e-90 | 1 | 425 | 12 | 439 | Acetyl esterase Axe7A OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe7A PE=1 SV=1 |
| P94388 | 6.04e-29 | 130 | 425 | 15 | 317 | Cephalosporin-C deacetylase OS=Bacillus subtilis (strain 168) OX=224308 GN=cah PE=1 SV=1 |
| Q9WXT2 | 1.71e-27 | 130 | 406 | 15 | 303 | Cephalosporin-C deacetylase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=axeA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
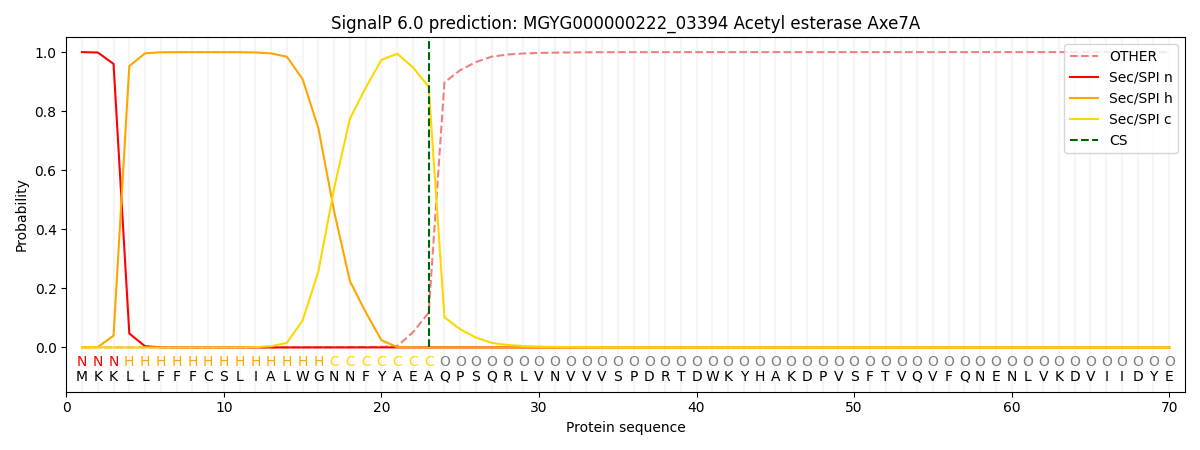
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000928 | 0.997946 | 0.000428 | 0.000247 | 0.000207 | 0.000197 |
