You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000226_01681
You are here: Home > Sequence: MGYG000000226_01681
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
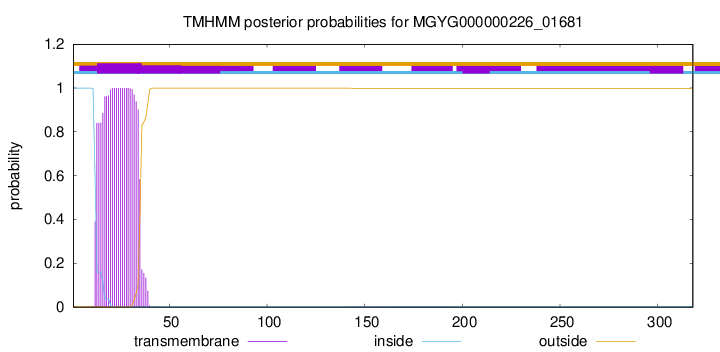
TMHMM annotations
Basic Information help
| Species | Lactococcus lactis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Lactococcus; Lactococcus lactis | |||||||||||
| CAZyme ID | MGYG000000226_01681 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 93168; End: 94124 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 136 | 265 | 1.7e-34 | 0.9307692307692308 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10918 | CE4_NodB_like_5s_6s | 9.53e-55 | 143 | 294 | 1 | 154 | Putative catalytic NodB homology domain of PgaB, IcaB, and similar proteins which consist of a deformed (beta/alpha)8 barrel fold with 5- or 6-strands. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes bacterial poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB, hemin storage system HmsF protein in gram-negative species, intercellular adhesion proteins IcaB, and many uncharacterized prokaryotic polysaccharide deacetylases. It also includes a putative polysaccharide deacetylase YxkH encoded by the Bacillus subtilis yxkH gene, which is one of six polysaccharide deacetylase gene homologs present in the Bacillus subtilis genome. Sequence comparison shows all family members contain a conserved domain similar to the catalytic NodB homology domain of rhizobial NodB-like proteins, which consists of a deformed (beta/alpha)8 barrel fold with 6 or 7 strands. However, in this family, most proteins have 5 strands and some have 6 strands. Moreover, long insertions are found in many family members, whose function remains unknown. |
| pfam01522 | Polysacc_deac_1 | 5.78e-44 | 136 | 266 | 1 | 124 | Polysaccharide deacetylase. This domain is found in polysaccharide deacetylase. This family of polysaccharide deacetylases includes NodB (nodulation protein B from Rhizobium) which is a chitooligosaccharide deacetylase. It also includes chitin deacetylase from yeast, and endoxylanases which hydrolyzes glucosidic bonds in xylan. |
| cd10969 | CE4_Ecf1_like_5s | 3.91e-43 | 108 | 292 | 3 | 200 | Putative catalytic NodB homology domain of a hypothetical protein Ecf1 from Escherichia coli and similar proteins. This family contains a hypothetical protein Ecf1 from Escherichia coli and its prokaryotic homologs. Although their biochemical properties remain to be determined, members in this family contain a conserved domain with a 5-stranded beta/alpha barrel, which is similar to the catalytic NodB homology domain of rhizobial NodB-like proteins, belonging to the larger carbohydrate esterase 4 (CE4) superfamily. |
| COG0726 | CDA1 | 5.04e-37 | 82 | 264 | 1 | 181 | Peptidoglycan/xylan/chitin deacetylase, PgdA/CDA1 family [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| cd10970 | CE4_DAC_u1_6s | 1.40e-34 | 143 | 307 | 2 | 158 | Putative catalytic NodB homology domain of uncharacterized prokaryotic polysaccharide deacetylases which consist of a 6-stranded beta/alpha barrel. This family contains uncharacterized prokaryotic polysaccharide deacetylases. Although their biological functions remain unknown, all members of the family contain a conserved domain with a 6-stranded beta/alpha barrel, which is similar to the catalytic NodB homology domain of rhizobial NodB-like proteins, belonging to the larger carbohydrate esterase 4 (CE4) superfamily. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QYR62058.1 | 2.86e-34 | 79 | 267 | 10 | 206 |
| QZY32711.1 | 8.72e-33 | 67 | 294 | 50 | 262 |
| QYR65360.1 | 1.17e-32 | 79 | 267 | 10 | 206 |
| QYR68238.1 | 4.50e-32 | 79 | 267 | 10 | 206 |
| QYR55309.1 | 2.42e-31 | 79 | 267 | 10 | 206 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6DQ3_A | 9.03e-51 | 81 | 309 | 8 | 230 | ChainA, Polysaccharide deacetylase [Streptococcus pyogenes],6DQ3_B Chain B, Polysaccharide deacetylase [Streptococcus pyogenes] |
| 4V33_A | 2.32e-15 | 77 | 307 | 139 | 359 | Crystalstructure of the putative polysaccharide deacetylase BA0330 from bacillus anthracis [Bacillus anthracis],4V33_B Crystal structure of the putative polysaccharide deacetylase BA0330 from bacillus anthracis [Bacillus anthracis] |
| 4HD5_A | 5.76e-15 | 77 | 307 | 139 | 359 | CrystalStructure of BC0361, a polysaccharide deacetylase from Bacillus cereus [Bacillus cereus ATCC 14579] |
| 4WCJ_A | 8.00e-15 | 82 | 263 | 37 | 224 | Structureof IcaB from Ammonifex degensii [Ammonifex degensii KC4] |
| 5BU6_A | 6.60e-11 | 144 | 273 | 78 | 249 | Structureof BpsB deaceylase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50],5BU6_B Structure of BpsB deaceylase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94361 | 5.27e-37 | 73 | 294 | 56 | 262 | Putative polysaccharide deacetylase YxkH OS=Bacillus subtilis (strain 168) OX=224308 GN=yxkH PE=3 SV=1 |
| Q6GDD6 | 1.17e-16 | 109 | 266 | 82 | 249 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=icaB PE=3 SV=1 |
| Q6G606 | 1.17e-16 | 109 | 266 | 82 | 249 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain MSSA476) OX=282459 GN=icaB PE=3 SV=1 |
| Q7A349 | 1.17e-16 | 109 | 266 | 82 | 249 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain N315) OX=158879 GN=icaB PE=3 SV=1 |
| Q8NUI6 | 1.17e-16 | 109 | 266 | 82 | 249 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain MW2) OX=196620 GN=icaB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000022 | 0.000003 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |