You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000227_00827
You are here: Home > Sequence: MGYG000000227_00827
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
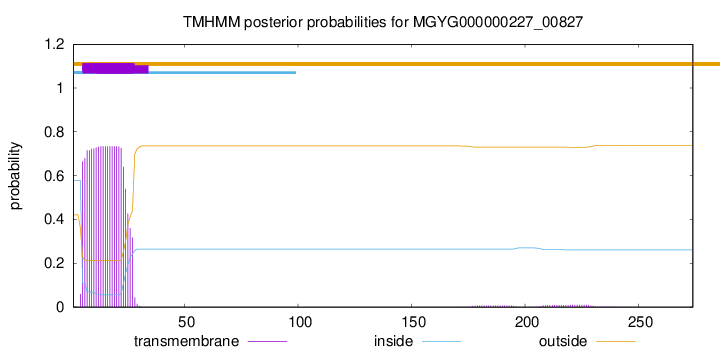
TMHMM annotations
Basic Information help
| Species | Bacillus sonorensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales; Bacillaceae; Bacillus; Bacillus sonorensis | |||||||||||
| CAZyme ID | MGYG000000227_00827 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | putative peptidoglycan endopeptidase LytE | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 821601; End: 822425 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00877 | NLPC_P60 | 5.75e-40 | 172 | 273 | 1 | 105 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| COG0791 | Spr | 1.85e-32 | 152 | 272 | 67 | 197 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
| PRK13914 | PRK13914 | 3.13e-25 | 24 | 273 | 198 | 480 | invasion associated endopeptidase. |
| PRK10838 | spr | 3.57e-20 | 161 | 273 | 68 | 183 | bifunctional murein DD-endopeptidase/murein LD-carboxypeptidase. |
| PRK06347 | PRK06347 | 4.30e-20 | 15 | 168 | 320 | 480 | 1,4-beta-N-acetylmuramoylhydrolase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ASB90330.1 | 4.54e-177 | 1 | 274 | 1 | 274 |
| ATH94870.1 | 3.04e-159 | 1 | 274 | 1 | 274 |
| SCA84862.1 | 4.31e-159 | 1 | 274 | 1 | 274 |
| QAT64418.1 | 4.31e-159 | 1 | 274 | 1 | 274 |
| QHZ46035.1 | 1.96e-145 | 1 | 274 | 1 | 271 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7CFL_A | 8.07e-19 | 158 | 273 | 12 | 136 | ChainA, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_B Chain B, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_C Chain C, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_D Chain D, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile] |
| 2K1G_A | 2.09e-16 | 172 | 273 | 18 | 122 | SolutionNMR structure of lipoprotein spr from Escherichia coli K12. Northeast Structural Genomics target ER541-37-162 [Escherichia coli K-12] |
| 4XCM_A | 9.44e-16 | 88 | 273 | 4 | 229 | Crystalstructure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4XCM_B Crystal structure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8] |
| 6B8C_A | 5.13e-12 | 158 | 253 | 26 | 123 | Crystalstructure of NlpC/p60 domain of peptidoglycan hydrolase SagA [Enterococcus faecium] |
| 3PVQ_A | 8.55e-09 | 154 | 252 | 150 | 259 | Crystalstructure of a putative dipeptidyl-peptidase VI (BT_1314) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482],3PVQ_B Crystal structure of a putative dipeptidyl-peptidase VI (BT_1314) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 2.10 A resolution [Bacteroides thetaiotaomicron VPI-5482],4R0K_A Crystal structure of a putative dipeptidyl-peptidase VI (BT_1314) from Bacteroides thetaiotaomicron VPI-5482 at 1.75 A resolution [Bacteroides thetaiotaomicron VPI-5482],4R0K_B Crystal structure of a putative dipeptidyl-peptidase VI (BT_1314) from Bacteroides thetaiotaomicron VPI-5482 at 1.75 A resolution [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P54421 | 1.75e-119 | 1 | 274 | 1 | 334 | Probable peptidoglycan endopeptidase LytE OS=Bacillus subtilis (strain 168) OX=224308 GN=lytE PE=1 SV=1 |
| O31852 | 7.58e-74 | 24 | 274 | 155 | 413 | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlS PE=1 SV=1 |
| O07532 | 3.68e-73 | 30 | 274 | 244 | 487 | Peptidoglycan endopeptidase LytF OS=Bacillus subtilis (strain 168) OX=224308 GN=lytF PE=1 SV=2 |
| Q01837 | 2.31e-30 | 24 | 273 | 194 | 523 | Probable endopeptidase p60 OS=Listeria ivanovii OX=1638 GN=iap PE=3 SV=1 |
| Q01838 | 1.42e-28 | 24 | 273 | 196 | 522 | Probable endopeptidase p60 OS=Listeria seeligeri OX=1640 GN=iap PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
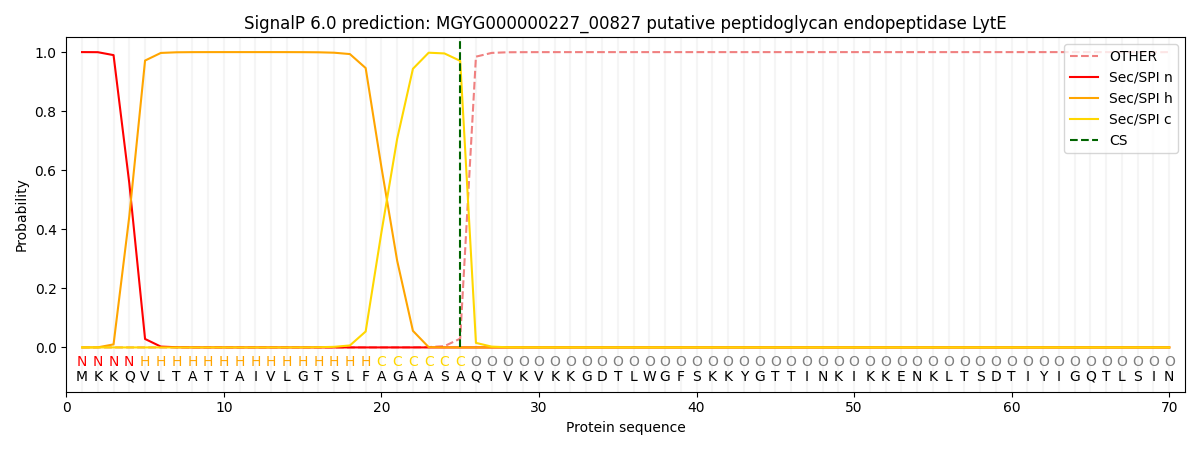
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000325 | 0.998886 | 0.000209 | 0.000221 | 0.000182 | 0.000160 |