You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000247_00946
You are here: Home > Sequence: MGYG000000247_00946
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
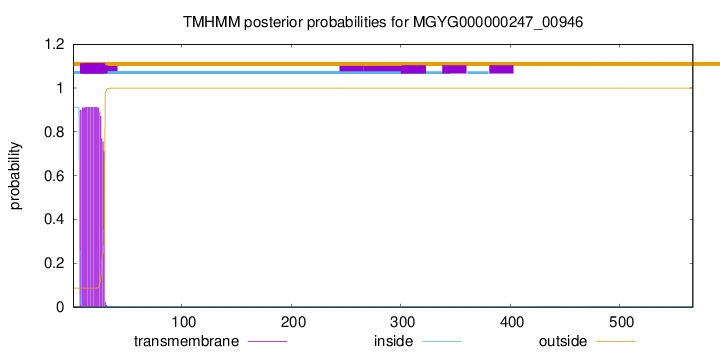
TMHMM annotations
Basic Information help
| Species | Collinsella intestinalis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; Collinsella intestinalis | |||||||||||
| CAZyme ID | MGYG000000247_00946 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1074304; End: 1076007 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 223 | 533 | 1.8e-43 | 0.5672514619883041 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 1.82e-29 | 224 | 539 | 3 | 204 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| COG4409 | NanH | 4.24e-07 | 229 | 280 | 269 | 321 | Neuraminidase (sialidase) [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| pfam13088 | BNR_2 | 1.23e-06 | 438 | 529 | 78 | 158 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| pfam13088 | BNR_2 | 0.009 | 218 | 294 | 147 | 216 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APO34972.1 | 2.71e-107 | 1 | 334 | 1 | 348 |
| ANU47084.1 | 7.06e-90 | 217 | 538 | 1204 | 1530 |
| QQQ98205.1 | 7.06e-90 | 217 | 538 | 1204 | 1530 |
| ARE59829.1 | 4.60e-50 | 218 | 524 | 35 | 322 |
| QQR06511.1 | 4.60e-50 | 218 | 524 | 35 | 322 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4YW1_A | 2.79e-24 | 206 | 522 | 177 | 489 | ChainA, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW1_B Chain B, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW2_A Chain A, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW2_B Chain B, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW3_A Chain A, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW3_B Chain B, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW4_A Streptococcus pneumoniae sialidase NanC [Streptococcus pneumoniae],4YW4_B Streptococcus pneumoniae sialidase NanC [Streptococcus pneumoniae],4YW5_A Chain A, Neuraminidase C [Streptococcus pneumoniae TIGR4],4YW5_B Chain B, Neuraminidase C [Streptococcus pneumoniae TIGR4],5F9T_A Chain A, Neuraminidase C [Streptococcus pneumoniae TIGR4],5F9T_B Chain B, Neuraminidase C [Streptococcus pneumoniae TIGR4] |
| 4YZ1_A | 2.91e-24 | 206 | 522 | 196 | 508 | CrystalStructure of Streptococcus pneumoniae NanC, apo structure. [Streptococcus pneumoniae TIGR4],4YZ1_B Crystal Structure of Streptococcus pneumoniae NanC, apo structure. [Streptococcus pneumoniae TIGR4],4YZ2_A Crystal Structure of Streptococcus pneumoniae NanC, in complex with 2-deoxy-2,3-didehydro-N-acetylneuraminic acid. [Streptococcus pneumoniae],4YZ2_B Crystal Structure of Streptococcus pneumoniae NanC, in complex with 2-deoxy-2,3-didehydro-N-acetylneuraminic acid. [Streptococcus pneumoniae],4YZ3_A Crystal Structure of Streptococcus pneumoniae NanC, in complex with Oseltamivir. [Streptococcus pneumoniae TIGR4],4YZ3_B Crystal Structure of Streptococcus pneumoniae NanC, in complex with Oseltamivir. [Streptococcus pneumoniae TIGR4],4YZ4_A Crystal Structure of Streptococcus pneumoniae NanC, in complex with N-Acetylneuraminic acid. [Streptococcus pneumoniae],4YZ4_B Crystal Structure of Streptococcus pneumoniae NanC, in complex with N-Acetylneuraminic acid. [Streptococcus pneumoniae],4YZ5_A Crystal Structure of Streptococcus pneumoniae NanC, in complex with 3-Sialyllactose [Streptococcus pneumoniae],4YZ5_B Crystal Structure of Streptococcus pneumoniae NanC, in complex with 3-Sialyllactose [Streptococcus pneumoniae] |
| 4X6K_A | 5.42e-24 | 225 | 521 | 9 | 307 | Crystalstructure of the intramolecular trans-sialidase from Ruminococcus gnavus in complex with Siastatin B [[Ruminococcus] gnavus CC55_001C] |
| 4X47_A | 5.54e-24 | 225 | 521 | 12 | 310 | Crystalstructure of the intramolecular trans-sialidase from Ruminococcus gnavus in complex with Neu5Ac2en [[Ruminococcus] gnavus ATCC 29149],4X49_A Crystal structure of the intramolecular trans-sialidase from Ruminococcus gnavus in complex with oseltamivir carboxylate [[Ruminococcus] gnavus ATCC 29149],4X4A_A Crystal structure of the intramolecular trans-sialidase from Ruminococcus gnavus in complex with 2,7-Anhydro-Neu5Ac [[Ruminococcus] gnavus ATCC 29149] |
| 5TSP_A | 2.70e-22 | 225 | 521 | 14 | 266 | Crystalstructure of the catalytic domain of Clostridium perfringens neuraminidase (NanI) in complex with a CHES [Clostridium perfringens ATCC 13124],5TSP_B Crystal structure of the catalytic domain of Clostridium perfringens neuraminidase (NanI) in complex with a CHES [Clostridium perfringens ATCC 13124] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P29767 | 8.14e-28 | 177 | 521 | 354 | 636 | Sialidase OS=Clostridium septicum OX=1504 PE=3 SV=1 |
| P62575 | 1.58e-20 | 206 | 521 | 317 | 616 | Sialidase A OS=Streptococcus pneumoniae OX=1313 GN=nanA PE=1 SV=1 |
| P62576 | 1.58e-20 | 206 | 521 | 317 | 616 | Sialidase A OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=nanA PE=1 SV=1 |
| Q27701 | 2.66e-18 | 217 | 521 | 271 | 580 | Anhydrosialidase OS=Macrobdella decora OX=6405 PE=1 SV=1 |
| P29768 | 1.18e-07 | 230 | 320 | 31 | 121 | Sialidase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=nanH PE=1 SV=4 |
SignalP and Lipop Annotations help
This protein is predicted as SP
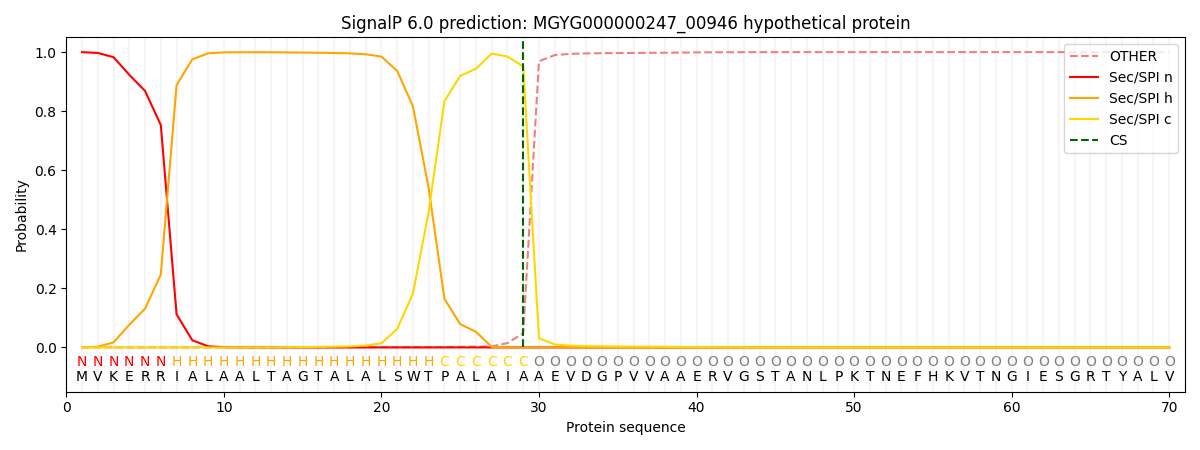
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000759 | 0.997936 | 0.000241 | 0.000597 | 0.000258 | 0.000193 |