You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000253_01604
You are here: Home > Sequence: MGYG000000253_01604
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
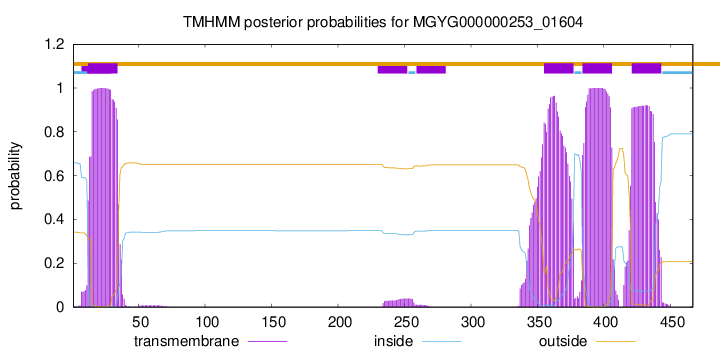
TMHMM annotations
Basic Information help
| Species | Holdemanella sp002299315 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelotrichaceae; Holdemanella; Holdemanella sp002299315 | |||||||||||
| CAZyme ID | MGYG000000253_01604 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13769; End: 15172 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 59 | 325 | 1.3e-27 | 0.9826086956521739 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06423 | CESA_like | 1.09e-63 | 61 | 276 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| COG1215 | BcsA | 4.37e-50 | 9 | 449 | 9 | 409 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| PRK11204 | PRK11204 | 1.02e-38 | 56 | 428 | 53 | 354 | N-glycosyltransferase; Provisional |
| pfam00535 | Glycos_transf_2 | 6.93e-28 | 60 | 235 | 1 | 152 | Glycosyl transferase family 2. Diverse family, transferring sugar from UDP-glucose, UDP-N-acetyl- galactosamine, GDP-mannose or CDP-abequose, to a range of substrates including cellulose, dolichol phosphate and teichoic acids. |
| PRK14583 | hmsR | 7.19e-27 | 59 | 336 | 77 | 319 | poly-beta-1,6 N-acetyl-D-glucosamine synthase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNK56534.1 | 5.18e-181 | 1 | 465 | 1 | 465 |
| BAN78238.1 | 6.96e-170 | 8 | 461 | 13 | 466 |
| QUO31269.1 | 2.72e-168 | 1 | 466 | 1 | 466 |
| CBL04658.1 | 6.71e-166 | 8 | 465 | 15 | 472 |
| QIB55150.1 | 6.00e-163 | 1 | 464 | 1 | 464 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5HEA_A | 1.37e-08 | 59 | 195 | 7 | 118 | CgTstructure in hexamer [Streptococcus parasanguinis FW213],5HEA_B CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEA_C CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEC_A CgT structure in dimer [Streptococcus parasanguinis FW213],5HEC_B CgT structure in dimer [Streptococcus parasanguinis FW213] |
| 5EKE_A | 5.32e-06 | 57 | 171 | 26 | 119 | Structureof the polyisoprenyl-phosphate glycosyltransferase GtrB (F215A mutant) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKE_B Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (F215A mutant) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKE_C Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (F215A mutant) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKE_D Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (F215A mutant) [Synechocystis sp. PCC 6803 substr. Kazusa] |
| 5EKP_A | 5.32e-06 | 57 | 171 | 26 | 119 | Structureof the polyisoprenyl-phosphate glycosyltransferase GtrB (WT) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKP_B Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (WT) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKP_C Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (WT) [Synechocystis sp. PCC 6803 substr. Kazusa],5EKP_D Structure of the polyisoprenyl-phosphate glycosyltransferase GtrB (WT) [Synechocystis sp. PCC 6803 substr. Kazusa] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5HCN1 | 1.43e-28 | 19 | 339 | 6 | 278 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain COL) OX=93062 GN=icaA PE=3 SV=1 |
| Q9RQP9 | 1.43e-28 | 19 | 339 | 6 | 278 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=icaA PE=3 SV=2 |
| Q7A351 | 1.43e-28 | 19 | 339 | 6 | 278 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain N315) OX=158879 GN=icaA PE=3 SV=1 |
| Q6GDD8 | 1.43e-28 | 19 | 339 | 6 | 278 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=icaA PE=3 SV=1 |
| Q99QX3 | 1.43e-28 | 19 | 339 | 6 | 278 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=icaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000045 | 0.000006 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |