You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000264_02881
You are here: Home > Sequence: MGYG000000264_02881
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
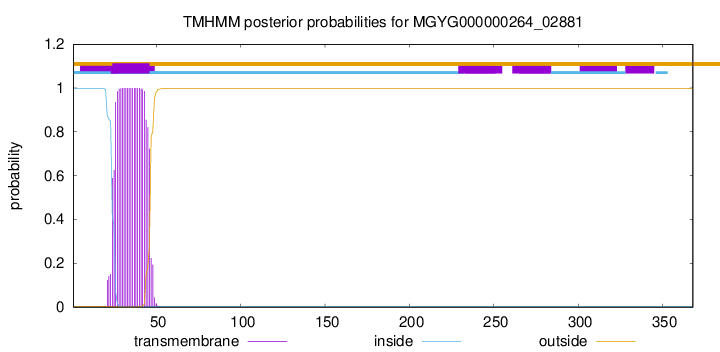
TMHMM annotations
Basic Information help
| Species | Exiguobacterium sp003467445 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Exiguobacterales; Exiguobacteraceae; Exiguobacterium; Exiguobacterium sp003467445 | |||||||||||
| CAZyme ID | MGYG000000264_02881 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13927; End: 15033 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16891 | CwlT-like | 2.97e-67 | 77 | 225 | 1 | 151 | CwlT-like N-terminal lysozyme domain and similar domains. CwlT is a bifunctional cell wall hydrolase containing an N-terminal lysozyme domain and a C-terminal NlpC/P60 endopeptidase domain (gamma-d-D-glutamyl-L-diamino acid endopeptidase), and has been implicated in the spread of transposons. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| pfam13702 | Lysozyme_like | 2.41e-58 | 71 | 224 | 1 | 165 | Lysozyme-like. |
| pfam01551 | Peptidase_M23 | 3.52e-35 | 268 | 362 | 3 | 96 | Peptidase family M23. Members of this family are zinc metallopeptidases with a range of specificities. The peptidase family M23 is included in this family, these are Gly-Gly endopeptidases. Peptidase family M23 are also endopeptidases. This family also includes some bacterial lipoproteins for which no proteolytic activity has been demonstrated. This family also includes leukocyte cell-derived chemotaxin 2 (LECT2) proteins. LECT2 is a liver-specific protein which is thought to be linked to hepatocyte growth although the exact function of this protein is unknown. |
| cd12797 | M23_peptidase | 1.11e-32 | 268 | 351 | 1 | 85 | M23 family metallopeptidase, also known as beta-lytic metallopeptidase, and similar proteins. This model describes the metallopeptidase M23 family, which includes beta-lytic metallopeptidase and lysostaphin. Members of this family are zinc endopeptidases that lyse bacterial cell wall peptidoglycans; they cleave either the N-acylmuramoyl-Ala bond between the cell wall peptidoglycan and the cross-linking peptide (e.g. beta-lytic endopeptidase) or a bond within the cross-linking peptide (e.g. stapholysin, and lysostaphin). Beta-lytic metallopeptidase, formerly known as beta-lytic protease, has a preference for cleavage of Gly-X bonds and favors hydrophobic or apolar residues on either side. It inhibits growth of sensitive organisms and may potentially serve as an antimicrobial agent. Lysostaphin, produced by Staphylococcus genus, cleaves pentaglycine cross-bridges of cell wall peptidoglycan, acting as autolysins to maintain cell wall metabolism or as toxins and weapons against competing strains. Staphylolysin (also known as LasA) is implicated in a range of processes related to Pseudomonas virulence, including stimulating shedding of the ectodomain of cell surface heparan sulphate proteoglycan syndecan-1, and elastin degradation in connective tissue. Its active site is less constricted and contains a five-coordinate zinc ion with trigonal bipyramidal geometry and two metal-bound water molecules, possibly contributing to its activity against a wider range of substrates than those used by related lytic enzymes, consistent with its multiple roles in Pseudomonas virulence. The family includes members that do not appear to have the conserved zinc-binding site and might be lipoproteins lacking proteolytic activity. |
| COG0739 | NlpD | 1.12e-32 | 240 | 364 | 128 | 262 | Murein DD-endopeptidase MepM and murein hydrolase activator NlpD, contain LysM domain [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CAQ35256.1 | 1.12e-247 | 1 | 368 | 1 | 368 |
| AHE40513.1 | 1.13e-246 | 1 | 368 | 1 | 364 |
| QUE87928.1 | 8.81e-245 | 1 | 368 | 1 | 368 |
| QUE87993.1 | 2.07e-243 | 1 | 368 | 1 | 368 |
| QUP88772.1 | 4.69e-242 | 1 | 366 | 1 | 366 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4FDY_A | 1.05e-19 | 66 | 224 | 18 | 178 | ChainA, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50],4FDY_B Chain B, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50] |
| 4HPE_A | 1.34e-19 | 67 | 360 | 17 | 300 | ChainA, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_B Chain B, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_C Chain C, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_D Chain D, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_E Chain E, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_F Chain F, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630] |
| 5KVP_A | 1.20e-17 | 262 | 366 | 28 | 152 | Solutionstructure of the catalytic domain of zoocin A [Streptococcus equi subsp. zooepidemicus] |
| 5J1L_A | 1.97e-16 | 245 | 352 | 40 | 152 | Crystalstructure of Csd1-Csd2 dimer I [Helicobacter pylori 26695],5J1L_C Crystal structure of Csd1-Csd2 dimer I [Helicobacter pylori 26695],5J1M_A Crystal structure of Csd1-Csd2 dimer II [Helicobacter pylori 26695],5J1M_C Crystal structure of Csd1-Csd2 dimer II [Helicobacter pylori 26695] |
| 3SLU_A | 5.78e-15 | 231 | 352 | 208 | 330 | Crystalstructure of NMB0315 [Neisseria meningitidis ATCC 13091],3SLU_B Crystal structure of NMB0315 [Neisseria meningitidis ATCC 13091] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96645 | 7.79e-34 | 59 | 224 | 45 | 192 | Probable endopeptidase YddH OS=Bacillus subtilis (strain 168) OX=224308 GN=yddH PE=3 SV=1 |
| O34636 | 1.60e-26 | 70 | 220 | 48 | 200 | Uncharacterized membrane protein YocA OS=Bacillus subtilis (strain 168) OX=224308 GN=yocA PE=4 SV=1 |
| Q99ZN9 | 2.00e-15 | 73 | 185 | 29 | 144 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M1 OX=301447 GN=pvaA PE=3 SV=1 |
| Q5XC63 | 2.74e-15 | 73 | 185 | 29 | 144 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M6 (strain ATCC BAA-946 / MGAS10394) OX=286636 GN=pvaA PE=3 SV=1 |
| Q8P120 | 2.74e-15 | 73 | 185 | 29 | 144 | Pneumococcal vaccine antigen A homolog OS=Streptococcus pyogenes serotype M18 (strain MGAS8232) OX=186103 GN=pvaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000015 | 0.000016 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |