You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000265_00044
You are here: Home > Sequence: MGYG000000265_00044
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides nordii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides nordii | |||||||||||
| CAZyme ID | MGYG000000265_00044 | |||||||||||
| CAZy Family | GT26 | |||||||||||
| CAZyme Description | Undecaprenyl-phosphate alpha-N-acetylglucosaminyl 1-phosphate transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 43366; End: 45204 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT26 | 428 | 581 | 1.7e-33 | 0.8771929824561403 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06853 | GT_WecA_like | 4.14e-66 | 49 | 315 | 1 | 249 | This subfamily contains Escherichia coli WecA, Bacillus subtilis TagO and related proteins. WecA is an UDP-N-acetylglucosamine (GlcNAc):undecaprenyl-phosphate (Und-P) GlcNAc-1-phosphate transferase that catalyzes the formation of a phosphodiester bond between a membrane-associated undecaprenyl-phosphate molecule and N-acetylglucosamine 1-phosphate, which is usually donated by a soluble UDP-N-acetylglucosamine precursor. WecA participates in the biosynthesis of O antigen LPS in many enteric bacteria and is also involved in the biosynthesis of enterobacterial common antigen. A conserved short sequence motif and a conserved arginine at a cytosolic loop of this integral membrane protein were shown to be critical in recognition of substrate UDP-N-acetylglucosamine. |
| pfam03808 | Glyco_tran_WecB | 4.13e-41 | 431 | 578 | 1 | 146 | Glycosyl transferase WecB/TagA/CpsF family. |
| COG0472 | Rfe | 2.02e-39 | 16 | 317 | 6 | 288 | UDP-N-acetylmuramyl pentapeptide phosphotransferase/UDP-N-acetylglucosamine-1-phosphate transferase [Cell wall/membrane/envelope biogenesis]. |
| cd06533 | Glyco_transf_WecG_TagA | 8.61e-38 | 428 | 578 | 1 | 148 | The glycosyltransferase WecG/TagA superfamily contains Escherichia coli WecG, Bacillus subtilis TagA and related proteins. E. coli WecG is believed to be a UDP-N-acetyl-D-mannosaminuronic acid transferase, and is involved in enterobacterial common antigen (eca) synthesis. B. subtilis TagA plays a key role in the Wall Teichoic Acid (WTA) biosynthetic pathway, catalyzing the transfer of N-acetylmannosamine to the C4 hydroxyl of a membrane-anchored N-acetylglucosaminyl diphospholipid to make ManNAc-beta-(1,4)-GlcNAc-pp-undecaprenyl. This is the first committed step in this pathway. Also included in this group is Xanthomonas campestris pv. campestris GumM, a glycosyltransferase participating in the biosynthesis of the exopolysaccharide xanthan. |
| cd06854 | GT_WbpL_WbcO_like | 1.62e-30 | 44 | 319 | 3 | 252 | The members of this subfamily catalyze the formation of a phosphodiester bond between a membrane-associated undecaprenyl-phosphate (Und-P) molecule and N-acetylhexosamine 1-phosphate, which is usually donated by a soluble UDP-N-acetylhexosamine precursor. The WbcO/WbpL substrate specificity has not yet been determined, but the structure of their biosynthetic end products implies that UDP-N-acetyl-D-fucosamine (UDP-FucNAc) and/or UDPN-acetyl-D-quinosamine (UDP-QuiNAc) are used. The subgroup of bacterial UDP-HexNAc:polyprenol-P HexNAc-1-P transferases includes the WbcO protein from Yersinia enterocolitica and the WbpL protein from Pseudomonas aeruginosa. These transferases initiate LPS O-antigen biosynthesis. Similar to other GlcNAc/MurNAc-1-P transferase family members, WbpL is a highly hydrophobic protein possessing 11 predicted transmembrane segments. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT75934.1 | 2.16e-257 | 45 | 612 | 1 | 589 |
| QJD98253.1 | 1.13e-51 | 403 | 576 | 17 | 190 |
| AMQ57519.1 | 2.06e-49 | 400 | 578 | 16 | 195 |
| QCT93965.1 | 6.07e-49 | 404 | 578 | 24 | 198 |
| APA64679.1 | 9.43e-49 | 394 | 579 | 19 | 206 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7MPK_A | 3.81e-18 | 406 | 580 | 34 | 206 | ChainA, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9],7MPK_B Chain B, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9],7MPK_C Chain C, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9],7N41_A Chain A, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9],7N41_B Chain B, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9],7N41_C Chain C, N-acetylglucosaminyldiphosphoundecaprenol N-acetyl-beta-D-mannosaminyltransferase [Thermoanaerobacter italicus Ab9] |
| 5WB4_A | 2.17e-17 | 386 | 570 | 23 | 195 | Crystalstructure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_B Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_C Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_D Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_E Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_F Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_G Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9],5WB4_H Crystal structure of the TarA wall teichoic acid glycosyltransferase [Thermoanaerobacter italicus Ab9] |
| 5WFG_A | 2.94e-17 | 386 | 570 | 23 | 195 | Crystalstructure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9],5WFG_B Crystal structure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9],5WFG_C Crystal structure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9],5WFG_D Crystal structure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9],5WFG_E Crystal structure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9],5WFG_F Crystal structure of the TarA wall teichoic acid glycosyltransferase bound to UDP [Thermoanaerobacter italicus Ab9] |
| 4J72_A | 2.26e-08 | 96 | 329 | 109 | 352 | CrystalStructure of polyprenyl-phosphate N-acetyl hexosamine 1-phosphate transferase [Aquifex aeolicus],4J72_B Crystal Structure of polyprenyl-phosphate N-acetyl hexosamine 1-phosphate transferase [Aquifex aeolicus],5CKR_A Crystal Structure of MraY in complex with Muraymycin D2 [Aquifex aeolicus VF5],6OYH_A Crystal structure of MraY bound to carbacaprazamycin [Aquifex aeolicus VF5],6OYH_B Crystal structure of MraY bound to carbacaprazamycin [Aquifex aeolicus VF5],6OYH_C Crystal structure of MraY bound to carbacaprazamycin [Aquifex aeolicus VF5],6OYH_D Crystal structure of MraY bound to carbacaprazamycin [Aquifex aeolicus VF5],6OYZ_A Crystal structure of MraY bound to capuramycin [Aquifex aeolicus VF5],6OYZ_B Crystal structure of MraY bound to capuramycin [Aquifex aeolicus VF5],6OYZ_C Crystal structure of MraY bound to capuramycin [Aquifex aeolicus VF5],6OYZ_D Crystal structure of MraY bound to capuramycin [Aquifex aeolicus VF5],6OZ6_A Crystal structure of MraY bound to 3'-hydroxymureidomycin A [Aquifex aeolicus VF5],6OZ6_B Crystal structure of MraY bound to 3'-hydroxymureidomycin A [Aquifex aeolicus VF5],6OZ6_C Crystal structure of MraY bound to 3'-hydroxymureidomycin A [Aquifex aeolicus VF5],6OZ6_D Crystal structure of MraY bound to 3'-hydroxymureidomycin A [Aquifex aeolicus VF5] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34753 | 1.87e-29 | 16 | 324 | 10 | 297 | Probable undecaprenyl-phosphate N-acetylglucosaminyl 1-phosphate transferase OS=Bacillus subtilis (strain 168) OX=224308 GN=tagO PE=1 SV=1 |
| Q8ZAE1 | 9.53e-27 | 42 | 326 | 33 | 300 | Undecaprenyl-phosphate alpha-N-acetylglucosaminyl 1-phosphate transferase OS=Yersinia pestis OX=632 GN=wecA PE=3 SV=1 |
| Q8XAS7 | 3.71e-25 | 36 | 337 | 27 | 311 | Undecaprenyl-phosphate alpha-N-acetylglucosaminyl 1-phosphate transferase OS=Escherichia coli O157:H7 OX=83334 GN=wecA PE=3 SV=1 |
| P0AC79 | 5.02e-25 | 36 | 337 | 27 | 311 | Undecaprenyl-phosphate alpha-N-acetylglucosaminyl 1-phosphate transferase OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=wecA PE=3 SV=1 |
| P0AC78 | 5.02e-25 | 36 | 337 | 27 | 311 | Undecaprenyl-phosphate alpha-N-acetylglucosaminyl 1-phosphate transferase OS=Escherichia coli (strain K12) OX=83333 GN=wecA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000053 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
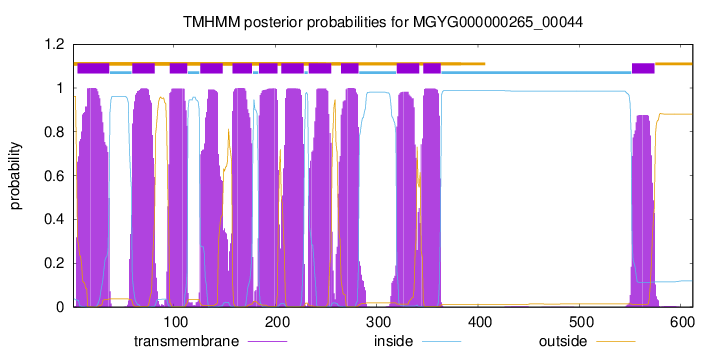
| start | end |
|---|---|
| 5 | 36 |
| 59 | 81 |
| 96 | 113 |
| 126 | 148 |
| 158 | 177 |
| 184 | 202 |
| 206 | 228 |
| 233 | 255 |
| 265 | 282 |
| 320 | 342 |
| 346 | 363 |
| 552 | 574 |
