You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000265_01936
You are here: Home > Sequence: MGYG000000265_01936
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides nordii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides nordii | |||||||||||
| CAZyme ID | MGYG000000265_01936 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 64199; End: 67081 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 555 | 840 | 5.6e-79 | 0.9858156028368794 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 9.05e-36 | 562 | 837 | 9 | 270 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.43e-19 | 557 | 877 | 54 | 402 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| TIGR04183 | Por_Secre_tail | 1.05e-04 | 907 | 959 | 21 | 71 | Por secretion system C-terminal sorting domain. Species that include Porphyromonas gingivalis, Fibrobacter succinogenes, Flavobacterium johnsoniae, Cytophaga hutchinsonii, Gramella forsetii, Prevotella intermedia, and Salinibacter ruber average twenty or more copies of a C-terminal domain, represented by this model, associated with sorting to the outer membrane and covalent modification. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI63368.1 | 6.92e-311 | 31 | 959 | 527 | 1466 |
| QNL39860.1 | 3.29e-94 | 543 | 865 | 27 | 356 |
| CBK67191.1 | 8.95e-94 | 543 | 865 | 27 | 356 |
| QDM10315.1 | 8.95e-94 | 543 | 865 | 27 | 356 |
| QGT72466.1 | 8.95e-94 | 543 | 865 | 27 | 356 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4EE9_A | 1.20e-67 | 556 | 873 | 5 | 313 | Crystalstructure of the RBcel1 endo-1,4-glucanase [uncultured bacterium],4M24_A Crystal structure of the endo-1,4-glucanase, RBcel1, in complex with cellobiose [uncultured bacterium] |
| 7P6G_A | 3.14e-67 | 556 | 873 | 5 | 313 | ChainA, Endoglucanase [uncultured bacterium],7P6G_B Chain B, Endoglucanase [uncultured bacterium],7P6H_A Chain A, Endoglucanase [uncultured bacterium],7P6H_B Chain B, Endoglucanase [uncultured bacterium] |
| 6ZZ3_A | 4.21e-67 | 556 | 873 | 5 | 313 | ChainA, Endoglucanase [uncultured bacterium],6ZZ3_B Chain B, Endoglucanase [uncultured bacterium],6ZZ3_C Chain C, Endoglucanase [uncultured bacterium],6ZZ3_D Chain D, Endoglucanase [uncultured bacterium] |
| 7P6I_A | 4.34e-67 | 556 | 873 | 5 | 313 | ChainA, Endoglucanase [uncultured bacterium],7P6J_A Chain A, Endoglucanase [uncultured bacterium],7P6J_B Chain B, Endoglucanase [uncultured bacterium],7P6J_C Chain C, Endoglucanase [uncultured bacterium],7P6J_D Chain D, Endoglucanase [uncultured bacterium] |
| 5LJF_A | 8.25e-67 | 556 | 873 | 5 | 313 | ChainA, Endoglucanase [uncultured bacterium],5LJF_B Chain B, Endoglucanase [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P58599 | 1.60e-37 | 557 | 867 | 119 | 424 | Endoglucanase OS=Ralstonia solanacearum (strain GMI1000) OX=267608 GN=egl PE=3 SV=1 |
| P17974 | 1.01e-36 | 557 | 867 | 121 | 426 | Endoglucanase OS=Ralstonia solanacearum OX=305 GN=egl PE=1 SV=2 |
| Q1HFS8 | 1.10e-26 | 555 | 848 | 29 | 310 | Endo-beta-1,4-glucanase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=eglB PE=1 SV=1 |
| B8MW97 | 2.47e-25 | 570 | 848 | 50 | 315 | probable endo-beta-1,4-glucanase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=eglB PE=3 SV=1 |
| Q2UPQ4 | 2.47e-25 | 570 | 848 | 50 | 315 | Probable endo-beta-1,4-glucanase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=eglB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
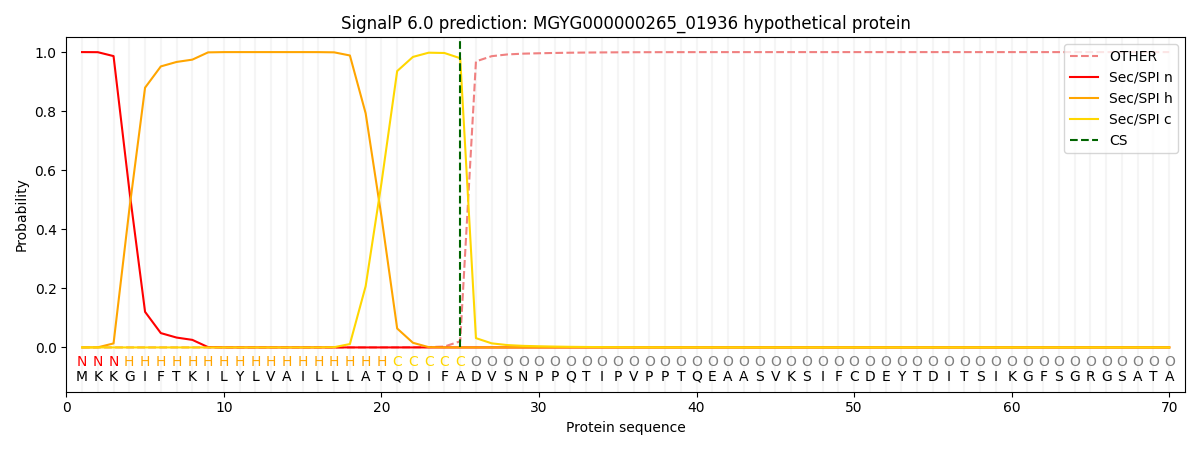
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000222 | 0.999195 | 0.000163 | 0.000147 | 0.000134 | 0.000128 |
