You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000265_02313
You are here: Home > Sequence: MGYG000000265_02313
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides nordii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides nordii | |||||||||||
| CAZyme ID | MGYG000000265_02313 | |||||||||||
| CAZy Family | GH97 | |||||||||||
| CAZyme Description | Retaining alpha-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 244859; End: 246787 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH97 | 11 | 641 | 6.6e-166 | 0.993660855784469 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam10566 | Glyco_hydro_97 | 4.42e-81 | 289 | 549 | 1 | 278 | Glycoside hydrolase 97. This domain is the catalytic region of the bacterial glycosyl-hydrolase family 97. This central part of the GH97 family protein sequences represents a typical and complete (beta/alpha)8-barrel or catalytic TIM-barrel type domain. The N- and C-terminal parts of the sequences, mainly consisting of beta-strands, form two additional non-catalytic domains. In all known glycosidases with the (beta-alpha)8-barrel fold, the amino acid residues at the active site are located on the C-termini of the beta-strands. |
| pfam14508 | GH97_N | 2.50e-52 | 26 | 284 | 1 | 235 | Glycosyl-hydrolase 97 N-terminal. This N-terminal domain of glycosyl-hydrolase-97 contributes part of the active site pocket. It is also important for contact with the catalytic and C-terminal domains of the whole. |
| pfam14509 | GH97_C | 9.19e-22 | 552 | 642 | 1 | 97 | Glycosyl-hydrolase 97 C-terminal, oligomerization. Glycosyl-hydrolase-97 is made up of three tightly linked and highly conserved globular domains. The C-terminal domain is found to be necessary for oligomerization of the whole molecule in order to create the active-site pocket and the Ca++-binding site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ58470.1 | 0.0 | 2 | 642 | 3 | 648 |
| QUT90417.1 | 0.0 | 2 | 642 | 3 | 648 |
| QDO71139.1 | 0.0 | 15 | 642 | 18 | 648 |
| ALJ60417.1 | 6.41e-214 | 17 | 640 | 31 | 654 |
| AYB33649.1 | 3.94e-207 | 4 | 637 | 11 | 637 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5XFM_A | 2.88e-93 | 49 | 636 | 40 | 637 | Crystalstructure of beta-arabinopyranosidase [Bacteroides thetaiotaomicron],5XFM_B Crystal structure of beta-arabinopyranosidase [Bacteroides thetaiotaomicron],5XFM_C Crystal structure of beta-arabinopyranosidase [Bacteroides thetaiotaomicron],5XFM_D Crystal structure of beta-arabinopyranosidase [Bacteroides thetaiotaomicron] |
| 3A24_A | 1.09e-64 | 26 | 642 | 6 | 641 | Crystalstructure of BT1871 retaining glycosidase [Bacteroides thetaiotaomicron],3A24_B Crystal structure of BT1871 retaining glycosidase [Bacteroides thetaiotaomicron] |
| 5E1Q_A | 1.37e-63 | 26 | 642 | 20 | 655 | Mutant(D415G) GH97 alpha-galactosidase in complex with Gal-Lac [Bacteroides thetaiotaomicron VPI-5482],5E1Q_B Mutant (D415G) GH97 alpha-galactosidase in complex with Gal-Lac [Bacteroides thetaiotaomicron VPI-5482] |
| 2D73_A | 7.66e-41 | 18 | 642 | 19 | 724 | CrystalStructure Analysis of SusB [Bacteroides thetaiotaomicron VPI-5482],2D73_B Crystal Structure Analysis of SusB [Bacteroides thetaiotaomicron VPI-5482],2ZQ0_A Crystal structure of SusB complexed with acarbose [Bacteroides thetaiotaomicron],2ZQ0_B Crystal structure of SusB complexed with acarbose [Bacteroides thetaiotaomicron] |
| 3WFA_A | 9.59e-41 | 24 | 642 | 5 | 704 | Catalyticrole of the calcium ion in GH97 inverting glycoside hydrolase [Bacteroides thetaiotaomicron],3WFA_B Catalytic role of the calcium ion in GH97 inverting glycoside hydrolase [Bacteroides thetaiotaomicron] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D7CFN7 | 5.94e-82 | 19 | 638 | 32 | 617 | Probable retaining alpha-galactosidase OS=Streptomyces bingchenggensis (strain BCW-1) OX=749414 GN=SBI_01652 PE=3 SV=1 |
| Q8A6L0 | 3.54e-70 | 2 | 642 | 3 | 662 | Retaining alpha-galactosidase OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_1871 PE=1 SV=1 |
| G8JZS4 | 1.39e-42 | 7 | 642 | 8 | 724 | Glucan 1,4-alpha-glucosidase SusB OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=susB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
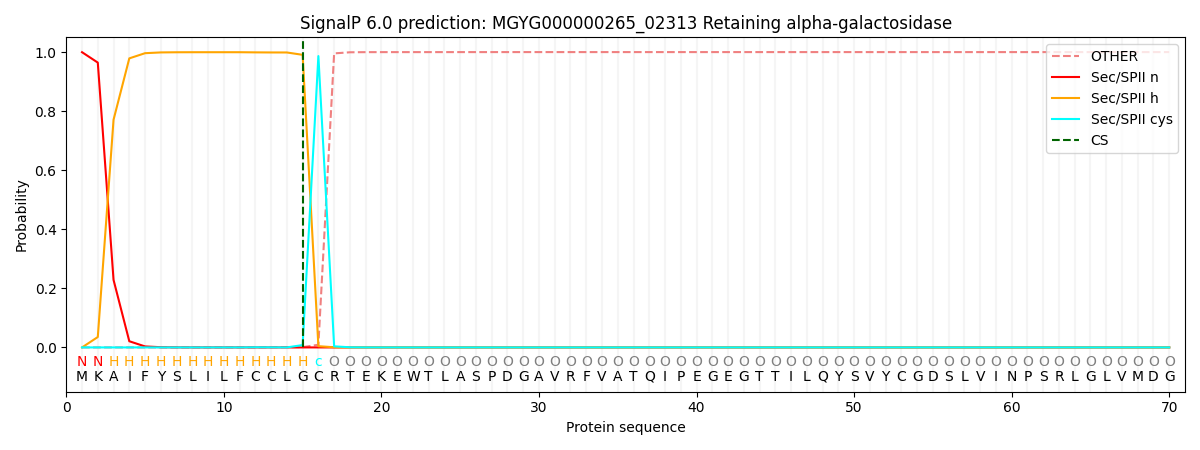
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000001 | 0.000313 | 0.999731 | 0.000000 | 0.000000 | 0.000000 |
