You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000265_03090
You are here: Home > Sequence: MGYG000000265_03090
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides nordii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides nordii | |||||||||||
| CAZyme ID | MGYG000000265_03090 | |||||||||||
| CAZy Family | GH37 | |||||||||||
| CAZyme Description | Cytoplasmic trehalase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 57324; End: 58652 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH37 | 24 | 434 | 1.8e-101 | 0.8370672097759674 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01204 | Trehalase | 1.01e-87 | 27 | 429 | 82 | 499 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| COG1626 | TreA | 1.86e-80 | 15 | 429 | 119 | 543 | Neutral trehalase [Carbohydrate transport and metabolism]. |
| PLN02567 | PLN02567 | 5.39e-67 | 50 | 429 | 132 | 535 | alpha,alpha-trehalase |
| PRK13270 | treF | 3.26e-61 | 53 | 429 | 154 | 537 | alpha,alpha-trehalase TreF. |
| PRK13272 | treA | 9.41e-59 | 36 | 429 | 124 | 527 | alpha,alpha-trehalase TreA. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI62569.1 | 1.30e-209 | 26 | 438 | 26 | 438 |
| QGA23972.1 | 1.87e-197 | 25 | 438 | 27 | 437 |
| BBL03724.1 | 1.23e-162 | 7 | 438 | 7 | 436 |
| BBL15914.1 | 4.97e-162 | 7 | 438 | 7 | 436 |
| QUB84795.1 | 1.70e-142 | 2 | 442 | 4 | 444 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5Z66_A | 6.91e-47 | 53 | 429 | 146 | 530 | Structureof periplasmic trehalase from Diamondback moth gut bacteria complexed with validoxylamine [Enterobacter cloacae],5Z6H_A Structure of periplasmic trehalase from Diamondback moth gut bacteria in the apo form [Enterobacter cloacae],5Z6H_B Structure of periplasmic trehalase from Diamondback moth gut bacteria in the apo form [Enterobacter cloacae] |
| 2JG0_A | 8.62e-47 | 30 | 429 | 83 | 493 | Family37 trehalase from Escherichia coli in complex with 1- thiatrehazolin [Escherichia coli K-12],2JJB_A Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_B Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_C Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_D Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2WYN_A Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_B Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_C Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_D Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12] |
| 2JF4_A | 2.17e-44 | 30 | 429 | 83 | 493 | Family37 trehalase from Escherichia coli in complex with validoxylamine [Escherichia coli K-12] |
| 5M4A_A | 6.41e-42 | 53 | 425 | 142 | 544 | Neutraltrehalase Nth1 from Saccharomyces cerevisiae in complex with trehalose [Saccharomyces cerevisiae] |
| 5NIS_A | 1.07e-41 | 53 | 425 | 195 | 597 | Neutraltrehalase Nth1 from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q31VA6 | 6.26e-51 | 53 | 429 | 154 | 537 | Cytoplasmic trehalase OS=Shigella boydii serotype 4 (strain Sb227) OX=300268 GN=treF PE=3 SV=1 |
| B7NEG2 | 1.21e-50 | 53 | 429 | 154 | 537 | Cytoplasmic trehalase OS=Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC) OX=585056 GN=treF PE=3 SV=1 |
| C4ZW66 | 1.21e-50 | 53 | 429 | 154 | 537 | Cytoplasmic trehalase OS=Escherichia coli (strain K12 / MC4100 / BW2952) OX=595496 GN=treF PE=3 SV=1 |
| B7UL72 | 1.21e-50 | 53 | 429 | 154 | 537 | Cytoplasmic trehalase OS=Escherichia coli O127:H6 (strain E2348/69 / EPEC) OX=574521 GN=treF PE=3 SV=1 |
| B7NNF3 | 1.21e-50 | 53 | 429 | 154 | 537 | Cytoplasmic trehalase OS=Escherichia coli O7:K1 (strain IAI39 / ExPEC) OX=585057 GN=treF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
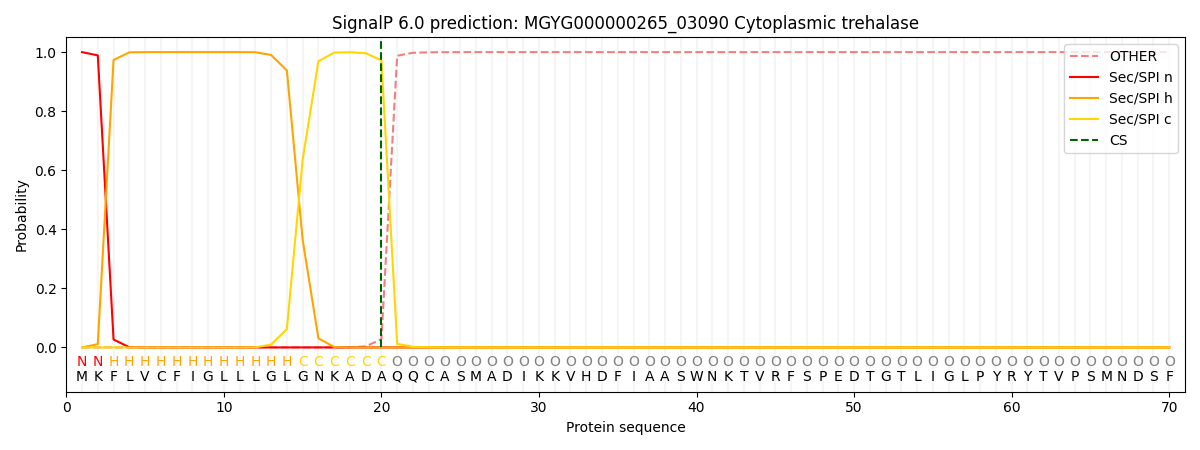
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000223 | 0.999162 | 0.000165 | 0.000142 | 0.000129 | 0.000127 |
