You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000268_01335
You are here: Home > Sequence: MGYG000000268_01335
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
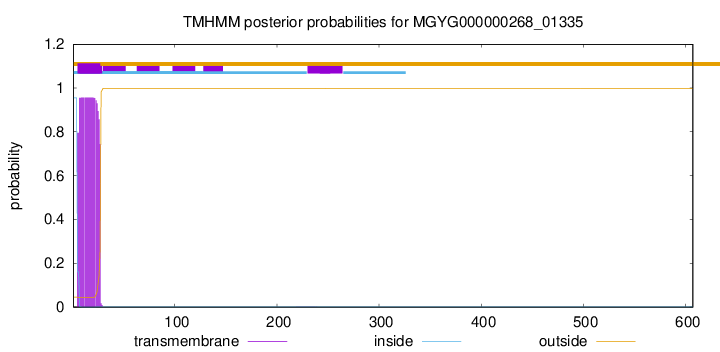
TMHMM annotations
Basic Information help
| Species | Schaedlerella sp900066545 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Schaedlerella; Schaedlerella sp900066545 | |||||||||||
| CAZyme ID | MGYG000000268_01335 | |||||||||||
| CAZy Family | GH25 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 82418; End: 84241 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH25 | 112 | 284 | 3.6e-44 | 0.9943502824858758 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06414 | GH25_LytC-like | 1.75e-93 | 109 | 292 | 1 | 189 | The LytC lysozyme of Streptococcus pneumoniae is a bacterial cell wall hydrolase that cleaves the beta1-4-glycosydic bond located between the N-acetylmuramoyl-N-glucosaminyl residues of the cell wall polysaccharide chains. LytC is composed of a C-terminal glycosyl hydrolase family 25 (GH25) domain and an N-terminal choline-binding module (CBM) consisting of eleven homologous repeats that specifically recognizes the choline residues of pneumococcal lipoteichoic and teichoic acids. This domain arrangement is the reverse of the major pneumococcal autolysin, LytA, and the CPL-1-like lytic enzymes of the pneumococcal bacteriophages, in which the CBM (consisting of six repeats) is at the C-terminus. This model represents the C-terminal catalytic domain of the LytC-like enzymes. |
| cd00599 | GH25_muramidase | 3.25e-38 | 110 | 292 | 1 | 185 | Endo-N-acetylmuramidases (muramidases) are lysozymes (also referred to as peptidoglycan hydrolases) that degrade bacterial cell walls by catalyzing the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues. This family of muramidases contains a glycosyl hydrolase family 25 (GH25) catalytic domain and is found in bacteria, fungi, slime molds, round worms, protozoans and bacteriophages. The bacteriophage members are referred to as endolysins which are involved in lysing the host cell at the end of the replication cycle to allow release of mature phage particles. Endolysins are typically modular enzymes consisting of a catalytically active domain that hydrolyzes the peptidoglycan cell wall and a cell wall-binding domain that anchors the protein to the cell wall. Endolysins generally have narrow substrate specificities with either intra-species or intra-genus bacteriolytic activity. |
| cd06525 | GH25_Lyc-like | 3.12e-25 | 110 | 292 | 1 | 183 | Lyc muramidase is an autolytic lysozyme (autolysin) from Clostridium acetobutylicum encoded by the lyc gene. Lyc has a glycosyl hydrolase family 25 (GH25) catalytic domain. Endo-N-acetylmuramidases are lysozymes (also referred to as peptidoglycan hydrolases) that degrade bacterial cell walls by catalyzing the hydrolysis of 1,4-beta-linkages between N-acetylmuramic acid and N-acetyl-D-glucosamine residues. |
| pfam01183 | Glyco_hydro_25 | 1.14e-22 | 112 | 284 | 1 | 180 | Glycosyl hydrolases family 25. |
| cd06524 | GH25_YegX-like | 2.82e-22 | 110 | 295 | 1 | 194 | YegX is an uncharacterized bacterial protein with a glycosyl hydrolase family 25 (GH25) catalytic domain that is similar in sequence to the CH-type (Chalaropsis-type) lysozymes of the GH25 family of endolysins. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWY97591.1 | 1.85e-95 | 72 | 603 | 41 | 588 |
| QIA33416.1 | 2.09e-80 | 94 | 598 | 11 | 528 |
| BAK44097.1 | 2.44e-74 | 36 | 304 | 527 | 834 |
| QCP36075.1 | 2.16e-72 | 64 | 294 | 35 | 286 |
| AZH69442.1 | 1.00e-69 | 76 | 292 | 35 | 263 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2WW5_A | 3.19e-24 | 74 | 291 | 234 | 465 | 3D-structureof the modular autolysin LytC from Streptococcus pneumoniae at 1.6 A resolution [Streptococcus pneumoniae R6],2WWD_A 3D-structure of the modular autolysin LytC from Streptococcus pneumoniae in complex with pneummococcal peptidoglycan fragment [Streptococcus pneumoniae R6] |
| 2WWC_A | 7.68e-24 | 74 | 291 | 234 | 465 | 3D-structureof the modular autolysin LytC from Streptococcus pneumoniae in complex with synthetic peptidoglycan ligand [Streptococcus pneumoniae R6] |
| 1JFX_A | 2.17e-11 | 110 | 298 | 6 | 207 | Crystalstructure of the bacterial lysozyme from Streptomyces coelicolor at 1.65 A resolution [Streptomyces coelicolor] |
| 4JZ5_A | 1.60e-09 | 112 | 291 | 26 | 210 | High-resolutionstructure of catalytic domain of endolysin ply40 from bacteriophage P40 of Listeria monocytogenes [Listeria phage P40] |
| 4KRU_A | 3.44e-08 | 110 | 291 | 21 | 205 | X-raystructure of catalytic domain of endolysin from clostridium perfringens phage phiSM101 [Clostridium phage phiSM101] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P25310 | 3.21e-10 | 110 | 298 | 83 | 284 | Lysozyme M1 OS=Streptomyces globisporus OX=1908 GN=acm PE=1 SV=1 |
| P76421 | 6.16e-10 | 108 | 309 | 66 | 265 | Uncharacterized protein YegX OS=Escherichia coli (strain K12) OX=83333 GN=yegX PE=3 SV=2 |
| Q8FFY2 | 1.11e-09 | 108 | 309 | 66 | 265 | Uncharacterized protein YegX OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=yegX PE=3 SV=2 |
| P26836 | 7.85e-06 | 110 | 291 | 10 | 194 | Probable autolytic lysozyme OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=lyc PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
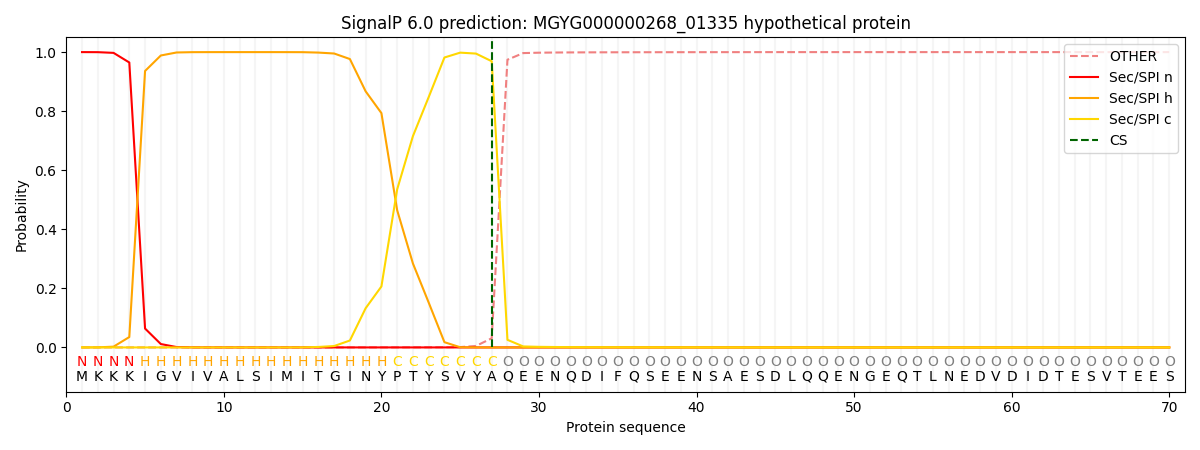
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000628 | 0.998387 | 0.000364 | 0.000202 | 0.000208 | 0.000177 |