You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000272_00572
You are here: Home > Sequence: MGYG000000272_00572
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella hominis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella hominis | |||||||||||
| CAZyme ID | MGYG000000272_00572 | |||||||||||
| CAZy Family | GH35 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10805; End: 13171 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH35 | 39 | 351 | 5.3e-121 | 0.993485342019544 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01301 | Glyco_hydro_35 | 6.90e-167 | 38 | 351 | 1 | 316 | Glycosyl hydrolases family 35. |
| PLN03059 | PLN03059 | 1.64e-50 | 31 | 608 | 29 | 720 | beta-galactosidase; Provisional |
| COG1874 | GanA | 6.78e-45 | 37 | 606 | 6 | 593 | Beta-galactosidase GanA [Carbohydrate transport and metabolism]. |
| pfam02449 | Glyco_hydro_42 | 8.15e-11 | 55 | 188 | 3 | 139 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| pfam13364 | BetaGal_dom4_5 | 8.52e-05 | 533 | 601 | 29 | 104 | Beta-galactosidase jelly roll domain. This domain is found in beta galactosidase enzymes. It has a jelly roll fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNT66372.1 | 0.0 | 1 | 788 | 1 | 788 |
| BCS84536.1 | 0.0 | 1 | 786 | 1 | 778 |
| AGB27610.1 | 0.0 | 1 | 786 | 1 | 796 |
| QVJ81893.1 | 0.0 | 30 | 786 | 22 | 786 |
| ADE83089.1 | 0.0 | 30 | 786 | 22 | 786 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6EON_A | 5.25e-306 | 30 | 787 | 26 | 773 | GalactanaseBT0290 [Bacteroides thetaiotaomicron VPI-5482] |
| 3D3A_A | 1.59e-252 | 30 | 606 | 6 | 576 | Crystalstructure of a beta-galactosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
| 4MAD_A | 3.48e-138 | 34 | 603 | 19 | 578 | ChainA, Beta-galactosidase [Niallia circulans],4MAD_B Chain B, Beta-galactosidase [Niallia circulans] |
| 7KDV_A | 1.18e-113 | 39 | 603 | 25 | 606 | ChainA, Beta-galactosidase [Mus musculus],7KDV_C Chain C, Beta-galactosidase [Mus musculus],7KDV_E Chain E, Beta-galactosidase [Mus musculus],7KDV_G Chain G, Beta-galactosidase [Mus musculus],7KDV_I Chain I, Beta-galactosidase [Mus musculus],7KDV_K Chain K, Beta-galactosidase [Mus musculus] |
| 3WF3_A | 3.11e-113 | 38 | 603 | 41 | 622 | Crystalstructure of human beta-galactosidase mutant I51T in complex with Galactose [Homo sapiens],3WF3_B Crystal structure of human beta-galactosidase mutant I51T in complex with Galactose [Homo sapiens],3WF3_C Crystal structure of human beta-galactosidase mutant I51T in complex with Galactose [Homo sapiens],3WF3_D Crystal structure of human beta-galactosidase mutant I51T in complex with Galactose [Homo sapiens],3WF4_A Crystal structure of human beta-galactosidase mutant I51T in complex with 6S-NBI-DGJ [Homo sapiens],3WF4_B Crystal structure of human beta-galactosidase mutant I51T in complex with 6S-NBI-DGJ [Homo sapiens],3WF4_C Crystal structure of human beta-galactosidase mutant I51T in complex with 6S-NBI-DGJ [Homo sapiens],3WF4_D Crystal structure of human beta-galactosidase mutant I51T in complex with 6S-NBI-DGJ [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48982 | 2.47e-130 | 3 | 586 | 2 | 573 | Beta-galactosidase OS=Xanthomonas manihotis OX=43353 GN=bga PE=1 SV=1 |
| P23780 | 3.73e-113 | 39 | 603 | 42 | 623 | Beta-galactosidase OS=Mus musculus OX=10090 GN=Glb1 PE=1 SV=1 |
| Q58D55 | 1.21e-112 | 39 | 613 | 40 | 630 | Beta-galactosidase OS=Bos taurus OX=9913 GN=GLB1 PE=2 SV=1 |
| P16278 | 1.66e-112 | 15 | 603 | 15 | 621 | Beta-galactosidase OS=Homo sapiens OX=9606 GN=GLB1 PE=1 SV=2 |
| Q5R7P4 | 1.77e-111 | 12 | 587 | 12 | 604 | Beta-galactosidase OS=Pongo abelii OX=9601 GN=GLB1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
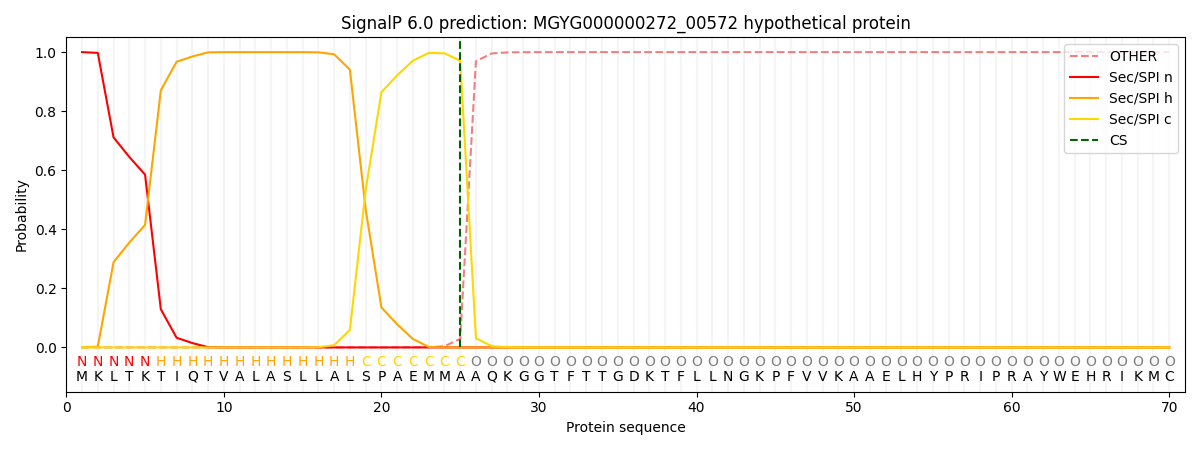
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000355 | 0.998732 | 0.000259 | 0.000246 | 0.000203 | 0.000167 |
