You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000272_01341
You are here: Home > Sequence: MGYG000000272_01341
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
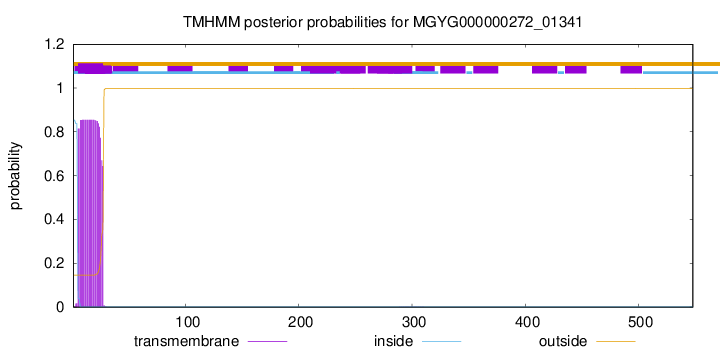
TMHMM annotations
Basic Information help
| Species | Prevotella hominis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella hominis | |||||||||||
| CAZyme ID | MGYG000000272_01341 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 53096; End: 54742 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 17 | 433 | 4.3e-102 | 0.9826589595375722 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00812 | Alpha_L_fucos | 6.45e-123 | 33 | 497 | 1 | 384 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| pfam01120 | Alpha_L_fucos | 9.18e-107 | 37 | 429 | 4 | 333 | Alpha-L-fucosidase. |
| COG3669 | AfuC | 6.75e-91 | 66 | 548 | 1 | 426 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
| pfam16757 | Fucosidase_C | 3.11e-04 | 474 | 534 | 15 | 73 | Alpha-L-fucosidase C-terminal domain. The C-terminal domain of Structure 1hl8 is constructed of eight anti-parallel-strands packed into two-sheets of five and three strands, respectively, forming a two-layer-sandwich containing a Greek key motif. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDO68463.1 | 0.0 | 20 | 548 | 19 | 546 |
| QUT88746.1 | 0.0 | 20 | 548 | 19 | 546 |
| ALJ60261.1 | 0.0 | 20 | 548 | 19 | 546 |
| QNL40547.1 | 0.0 | 16 | 548 | 17 | 548 |
| QUT56786.1 | 0.0 | 3 | 548 | 4 | 548 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7LJJ_A | 2.58e-86 | 35 | 547 | 37 | 509 | ChainA, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135],7LK7_A Chain A, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135] |
| 7LNP_A | 1.40e-85 | 35 | 547 | 37 | 509 | ChainA, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135],7LNP_C Chain C, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135],7LNP_D Chain D, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135],7LNP_E Chain E, Exo-alpha-L-galactosidase [Phocaeicola plebeius DSM 17135] |
| 4NI3_A | 2.54e-58 | 37 | 546 | 5 | 489 | ChainA, Alpha-fucosidase GH29 [Fusarium graminearum],4NI3_B Chain B, Alpha-fucosidase GH29 [Fusarium graminearum],4PSP_A Chain A, Alpha-fucosidase GH29 [Fusarium graminearum],4PSP_B Chain B, Alpha-fucosidase GH29 [Fusarium graminearum],4PSR_A Chain A, Alpha-fucosidase GH29 [Fusarium graminearum],4PSR_B Chain B, Alpha-fucosidase GH29 [Fusarium graminearum] |
| 1ODU_A | 4.65e-42 | 35 | 538 | 4 | 436 | CrystalStructure Of Thermotoga Maritima Alpha-Fucosidase In Complex With Fucose [Thermotoga maritima MSB8],1ODU_B Crystal Structure Of Thermotoga Maritima Alpha-Fucosidase In Complex With Fucose [Thermotoga maritima MSB8] |
| 2ZWY_A | 5.19e-42 | 35 | 538 | 4 | 436 | alpha-L-fucosidase[Thermotoga maritima],2ZWY_B alpha-L-fucosidase [Thermotoga maritima],2ZWZ_A alpha-L-fucosidase complexed with inhibitor, Core1 [Thermotoga maritima],2ZWZ_B alpha-L-fucosidase complexed with inhibitor, Core1 [Thermotoga maritima],2ZX5_A alpha-L-fucosidase complexed with inhibitor, F10 [Thermotoga maritima],2ZX5_B alpha-L-fucosidase complexed with inhibitor, F10 [Thermotoga maritima],2ZX6_A alpha-L-fucosidase complexed with inhibitor, F10-1C [Thermotoga maritima],2ZX6_B alpha-L-fucosidase complexed with inhibitor, F10-1C [Thermotoga maritima],2ZX7_A alpha-L-fucosidase complexed with inhibitor, F10-2C [Thermotoga maritima],2ZX7_B alpha-L-fucosidase complexed with inhibitor, F10-2C [Thermotoga maritima],2ZX8_A alpha-L-fucosidase complexed with inhibitor, F10-2C-O [Thermotoga maritima],2ZX8_B alpha-L-fucosidase complexed with inhibitor, F10-2C-O [Thermotoga maritima],2ZX9_A alpha-L-fucosidase complexed with inhibitor, B4 [Thermotoga maritima],2ZX9_B alpha-L-fucosidase complexed with inhibitor, B4 [Thermotoga maritima],2ZXA_A alpha-L-fucosidase complexed with inhibitor, FNJ-acetyl [Thermotoga maritima],2ZXA_B alpha-L-fucosidase complexed with inhibitor, FNJ-acetyl [Thermotoga maritima],2ZXB_A alpha-L-fucosidase complexed with inhibitor, ph-6FNJ [Thermotoga maritima],2ZXB_B alpha-L-fucosidase complexed with inhibitor, ph-6FNJ [Thermotoga maritima],2ZXD_A alpha-L-fucosidase complexed with inhibitor, iso-6FNJ [Thermotoga maritima],2ZXD_B alpha-L-fucosidase complexed with inhibitor, iso-6FNJ [Thermotoga maritima] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P17164 | 1.55e-33 | 39 | 534 | 33 | 442 | Tissue alpha-L-fucosidase OS=Rattus norvegicus OX=10116 GN=Fuca1 PE=1 SV=1 |
| P48300 | 1.94e-32 | 39 | 537 | 37 | 448 | Tissue alpha-L-fucosidase OS=Canis lupus familiaris OX=9615 GN=FUCA1 PE=2 SV=1 |
| Q9BTY2 | 6.85e-32 | 39 | 540 | 34 | 453 | Plasma alpha-L-fucosidase OS=Homo sapiens OX=9606 GN=FUCA2 PE=1 SV=2 |
| Q99LJ1 | 1.93e-31 | 39 | 534 | 23 | 432 | Tissue alpha-L-fucosidase OS=Mus musculus OX=10090 GN=Fuca1 PE=1 SV=1 |
| Q5RFI5 | 2.29e-31 | 39 | 540 | 32 | 451 | Plasma alpha-L-fucosidase OS=Pongo abelii OX=9601 GN=FUCA2 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000246 | 0.999027 | 0.000176 | 0.000174 | 0.000168 | 0.000161 |