You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000284_02453
Basic Information
help
| Species |
Parabacteroides sp900540715
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp900540715
|
| CAZyme ID |
MGYG000000284_02453
|
| CAZy Family |
GH106 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000284 |
5935659 |
MAG |
Sweden |
Europe |
|
| Gene Location |
Start: 55101;
End: 57281
Strand: -
|
No EC number prediction in MGYG000000284_02453.
| Family |
Start |
End |
Evalue |
family coverage |
| GH106 |
25 |
723 |
1.7e-82 |
0.7803398058252428 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam17132
|
Glyco_hydro_106 |
8.17e-33 |
156 |
699 |
338 |
873 |
alpha-L-rhamnosidase. |
This protein is predicted as SP
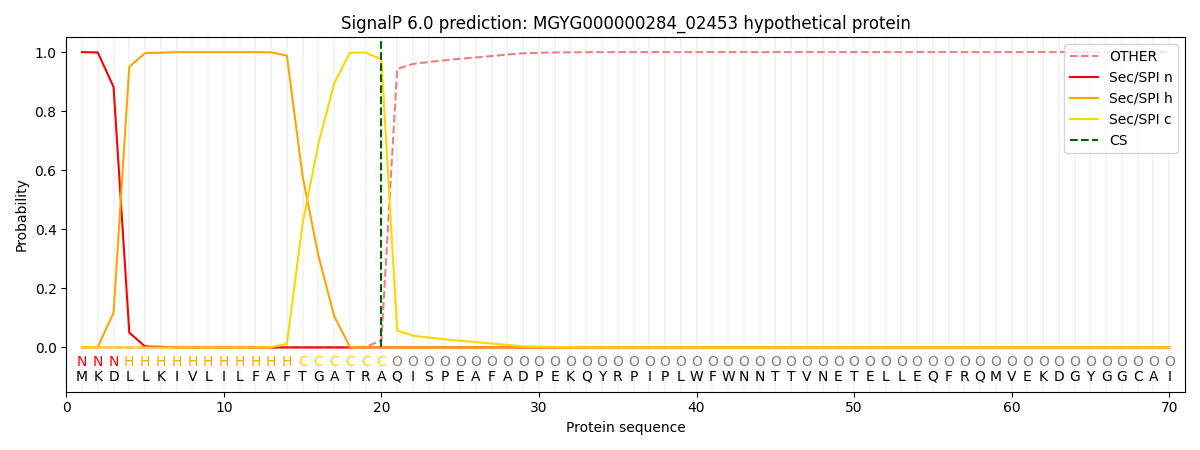
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000259
|
0.999090
|
0.000146
|
0.000170
|
0.000154
|
0.000145
|
There is no transmembrane helices in MGYG000000284_02453.

