You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000284_04638
You are here: Home > Sequence: MGYG000000284_04638
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides sp900540715 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp900540715 | |||||||||||
| CAZyme ID | MGYG000000284_04638 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2805; End: 5609 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 19 | 494 | 4.7e-73 | 0.5093085106382979 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 4.15e-29 | 23 | 564 | 12 | 541 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 1.36e-23 | 25 | 342 | 14 | 296 | beta-D-glucuronidase; Provisional |
| PRK10340 | ebgA | 2.58e-15 | 24 | 492 | 42 | 509 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam02837 | Glyco_hydro_2_N | 3.16e-14 | 24 | 216 | 2 | 169 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
| pfam00703 | Glyco_hydro_2 | 2.76e-12 | 219 | 322 | 2 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT75482.1 | 0.0 | 1 | 934 | 1 | 934 |
| QRO25183.1 | 0.0 | 14 | 931 | 16 | 930 |
| SCD20884.1 | 0.0 | 1 | 931 | 1 | 941 |
| ATP55310.1 | 0.0 | 6 | 932 | 12 | 952 |
| BBE19496.1 | 0.0 | 1 | 931 | 1 | 941 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7KGY_A | 6.99e-14 | 25 | 468 | 19 | 439 | ChainA, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_B Chain B, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_C Chain C, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_D Chain D, Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 6ED2_A | 2.89e-13 | 26 | 465 | 42 | 464 | ChainA, Glycosyl hydrolase family 2, TIM barrel domain protein [Faecalibacterium duncaniae] |
| 3K4A_A | 1.91e-12 | 24 | 342 | 15 | 294 | Crystalstructure of selenomethionine substituted E. coli beta-glucuronidase [Escherichia coli K-12],3K4A_B Crystal structure of selenomethionine substituted E. coli beta-glucuronidase [Escherichia coli K-12] |
| 4JHZ_A | 5.74e-12 | 24 | 342 | 15 | 294 | Structureof E. coli beta-Glucuronidase bound with a novel, potent inhibitor 2-[4-(1,3-benzodioxol-5-ylmethyl)piperazin-1-yl]-N-[(1S,2S,5S)-2,5-dimethoxycyclohexyl]acetamide [Escherichia coli K-12],4JHZ_B Structure of E. coli beta-Glucuronidase bound with a novel, potent inhibitor 2-[4-(1,3-benzodioxol-5-ylmethyl)piperazin-1-yl]-N-[(1S,2S,5S)-2,5-dimethoxycyclohexyl]acetamide [Escherichia coli K-12] |
| 3LPF_A | 5.76e-12 | 24 | 342 | 15 | 294 | Structureof E. coli beta-Glucuronidase bound with a novel, potent inhibitor 1-((6,7-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)-1-(2-hydroxyethyl)-3-(3-methoxyphenyl)thiourea [Escherichia coli K-12],3LPF_B Structure of E. coli beta-Glucuronidase bound with a novel, potent inhibitor 1-((6,7-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)-1-(2-hydroxyethyl)-3-(3-methoxyphenyl)thiourea [Escherichia coli K-12],3LPG_A Structure of E. coli beta-Glucuronidase bound with a novel, potent inhibitor 3-(2-fluorophenyl)-1-(2-hydroxyethyl)-1-((6-methyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)urea [Escherichia coli K-12],3LPG_B Structure of E. coli beta-Glucuronidase bound with a novel, potent inhibitor 3-(2-fluorophenyl)-1-(2-hydroxyethyl)-1-((6-methyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)urea [Escherichia coli K-12],5CZK_A Structure of E. coli beta-glucuronidase bound with a novel, potent inhibitor 1-((6,8-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)-1-(2-hydroxyethyl)-3-(4-hydroxyphenyl)thiourea [Escherichia coli K-12],5CZK_B Structure of E. coli beta-glucuronidase bound with a novel, potent inhibitor 1-((6,8-dimethyl-2-oxo-1,2-dihydroquinolin-3-yl)methyl)-1-(2-hydroxyethyl)-3-(4-hydroxyphenyl)thiourea [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A6TI29 | 7.93e-16 | 18 | 492 | 47 | 507 | Beta-galactosidase 2 OS=Klebsiella pneumoniae subsp. pneumoniae (strain ATCC 700721 / MGH 78578) OX=272620 GN=lacZ2 PE=3 SV=1 |
| A7KGA5 | 1.04e-15 | 18 | 492 | 47 | 507 | Beta-galactosidase OS=Klebsiella pneumoniae OX=573 GN=lacZ PE=3 SV=1 |
| Q0TKT1 | 3.10e-15 | 18 | 492 | 47 | 507 | Beta-galactosidase OS=Escherichia coli O6:K15:H31 (strain 536 / UPEC) OX=362663 GN=lacZ PE=3 SV=1 |
| Q1RFJ2 | 4.08e-15 | 18 | 492 | 47 | 507 | Beta-galactosidase OS=Escherichia coli (strain UTI89 / UPEC) OX=364106 GN=lacZ PE=3 SV=1 |
| A1A831 | 4.08e-15 | 18 | 492 | 47 | 507 | Beta-galactosidase OS=Escherichia coli O1:K1 / APEC OX=405955 GN=lacZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
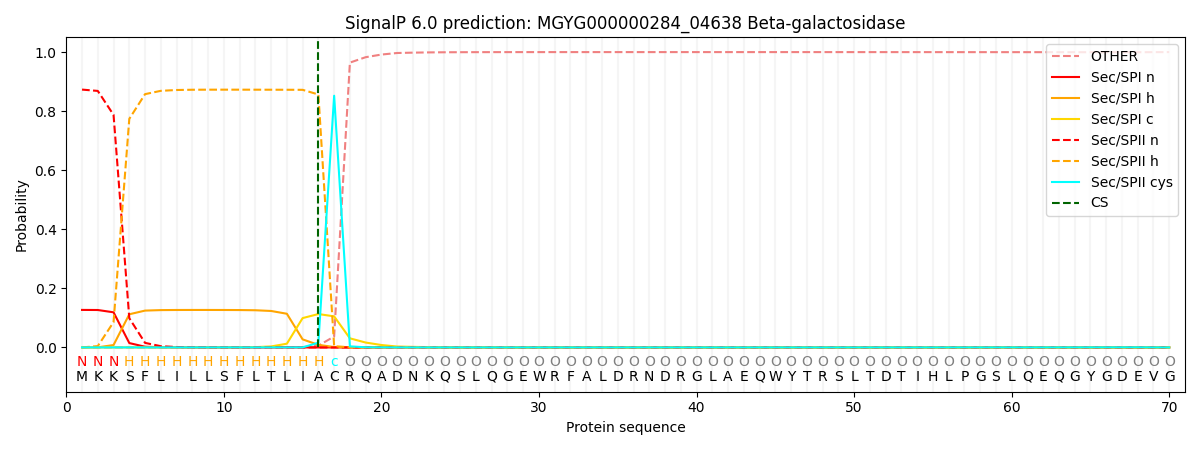
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000135 | 0.124490 | 0.875238 | 0.000048 | 0.000061 | 0.000051 |
