You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000295_01661
You are here: Home > Sequence: MGYG000000295_01661
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
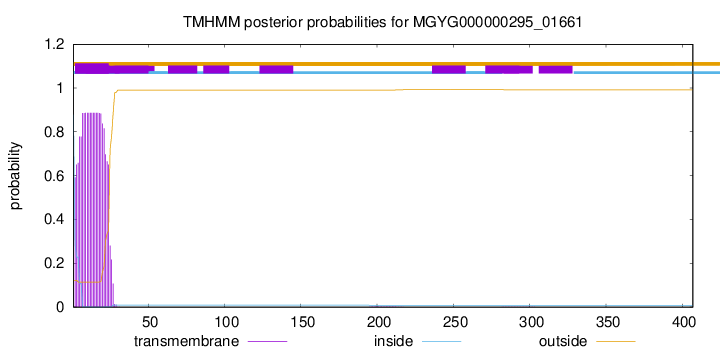
TMHMM annotations
Basic Information help
| Species | Clostridium_E sporosphaeroides | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; Clostridium_E; Clostridium_E sporosphaeroides | |||||||||||
| CAZyme ID | MGYG000000295_01661 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 631; End: 1854 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 142 | 364 | 1.7e-38 | 0.9583333333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00933 | Glyco_hydro_3 | 3.58e-55 | 85 | 397 | 1 | 314 | Glycosyl hydrolase family 3 N terminal domain. |
| COG1472 | BglX | 5.81e-47 | 84 | 407 | 1 | 319 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| PRK05337 | PRK05337 | 1.11e-26 | 127 | 364 | 39 | 278 | beta-hexosaminidase; Provisional |
| PRK15098 | PRK15098 | 9.38e-17 | 71 | 407 | 34 | 360 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNK41914.1 | 1.48e-130 | 54 | 407 | 61 | 420 |
| QAT49201.1 | 1.07e-128 | 68 | 407 | 84 | 423 |
| QHQ63470.1 | 9.85e-115 | 76 | 407 | 84 | 415 |
| BCJ98858.1 | 1.34e-108 | 76 | 407 | 127 | 458 |
| QWT55932.1 | 4.10e-108 | 81 | 407 | 112 | 438 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3BMX_A | 2.04e-31 | 77 | 405 | 35 | 398 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 3LK6_A | 8.37e-31 | 77 | 405 | 9 | 372 | ChainA, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
| 4GYJ_A | 9.54e-31 | 77 | 405 | 39 | 402 | Crystalstructure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYJ_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYK_A Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168],4GYK_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168] |
| 6K5J_A | 2.82e-29 | 84 | 405 | 11 | 342 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 4ZM6_A | 3.60e-29 | 85 | 400 | 8 | 337 | Aunique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432],4ZM6_B A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P40406 | 1.12e-30 | 77 | 405 | 35 | 398 | Beta-hexosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagZ PE=1 SV=1 |
| Q5H1Q0 | 2.13e-27 | 129 | 364 | 39 | 276 | Beta-hexosaminidase OS=Xanthomonas oryzae pv. oryzae (strain KACC10331 / KXO85) OX=291331 GN=nagZ PE=3 SV=2 |
| Q2P4L0 | 5.57e-27 | 129 | 364 | 39 | 276 | Beta-hexosaminidase OS=Xanthomonas oryzae pv. oryzae (strain MAFF 311018) OX=342109 GN=nagZ PE=3 SV=1 |
| Q7WUL3 | 7.20e-27 | 80 | 405 | 22 | 354 | Beta-N-acetylglucosaminidase/beta-glucosidase OS=Cellulomonas fimi OX=1708 GN=nag3 PE=1 SV=1 |
| Q8PMU1 | 1.45e-26 | 129 | 364 | 39 | 276 | Beta-hexosaminidase OS=Xanthomonas axonopodis pv. citri (strain 306) OX=190486 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
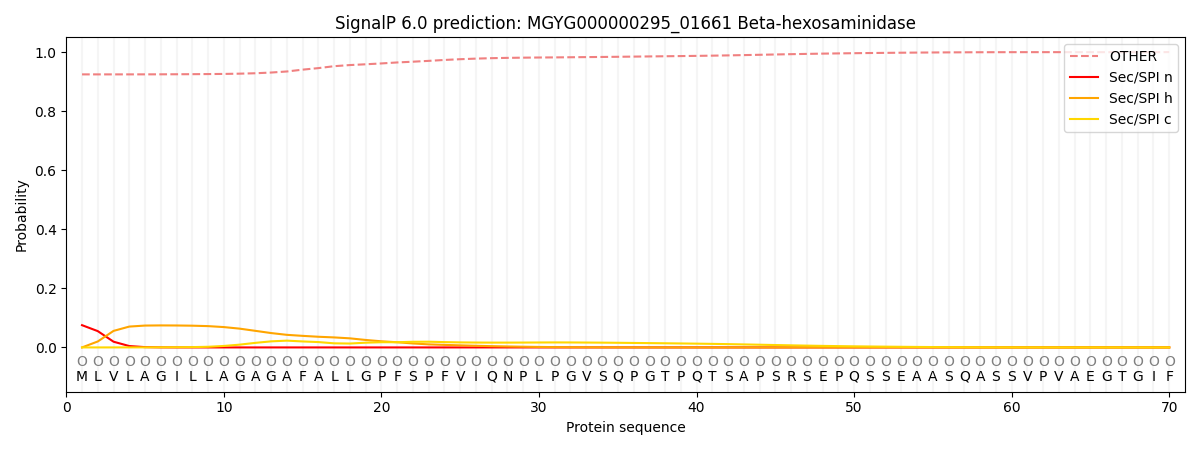
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.928675 | 0.070902 | 0.000199 | 0.000071 | 0.000057 | 0.000103 |