You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000298_01526
You are here: Home > Sequence: MGYG000000298_01526
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; | |||||||||||
| CAZyme ID | MGYG000000298_01526 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 254; End: 1528 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd03784 | GT1_Gtf-like | 3.48e-27 | 5 | 385 | 2 | 373 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| COG1819 | YjiC | 7.57e-25 | 3 | 424 | 1 | 406 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| pfam04101 | Glyco_tran_28_C | 0.002 | 256 | 351 | 2 | 101 | Glycosyltransferase family 28 C-terminal domain. The glycosyltransferase family 28 includes monogalactosyldiacylglycerol synthase (EC 2.4.1.46) and UDP-N-acetylglucosamine transferase (EC 2.4.1.-). Structural analysis suggests the C-terminal domain contains the UDP-GlcNAc binding site. |
| COG0707 | MurG | 0.010 | 301 | 403 | 225 | 338 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QLF56223.1 | 2.59e-279 | 1 | 424 | 1 | 424 |
| AMD97834.1 | 2.52e-270 | 1 | 424 | 1 | 424 |
| QRO08088.1 | 1.92e-239 | 1 | 422 | 1 | 422 |
| QQL00321.1 | 1.92e-239 | 1 | 422 | 1 | 422 |
| QLL97959.1 | 1.92e-239 | 1 | 422 | 1 | 422 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6KQW_A | 7.64e-13 | 1 | 365 | 1 | 330 | ChainA, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168] |
| 6KQX_A | 7.82e-13 | 1 | 365 | 1 | 330 | ChainA, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168],7BOV_A Chain A, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168] |
| 7LVY_A | 4.65e-07 | 254 | 385 | 296 | 424 | ChainA, UDP-glycosyltransferase 203A2 [Tetranychus urticae],7MCO_A Chain A, UDP-glycosyltransferase 203A2 [Tetranychus urticae],7MCO_B Chain B, UDP-glycosyltransferase 203A2 [Tetranychus urticae] |
| 3OTG_A | 5.65e-07 | 15 | 390 | 32 | 379 | CrystalStructure of CalG1, Calicheamicin Glycostyltransferase, TDP bound form [Micromonospora echinospora],3OTH_A Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora],3OTH_B Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora] |
| 4G2T_A | 6.97e-06 | 98 | 380 | 117 | 374 | CrystalStructure of Streptomyces sp. SF2575 glycosyltransferase SsfS6, complexed with thymidine diphosphate [Streptomyces sp. SF2575] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34539 | 4.28e-12 | 1 | 365 | 1 | 330 | NDP-glycosyltransferase YjiC OS=Bacillus subtilis (strain 168) OX=224308 GN=yjiC PE=1 SV=1 |
| P0DTE5 | 7.27e-08 | 75 | 355 | 111 | 408 | UDP-glucuronosyltransferase 2A2 OS=Homo sapiens OX=9606 GN=UGT2A2 PE=1 SV=1 |
| Q6PDD0 | 1.67e-07 | 98 | 355 | 136 | 400 | UDP-glucuronosyltransferase 2A2 OS=Mus musculus OX=10090 GN=Ugt2a2 PE=2 SV=1 |
| P36512 | 1.59e-06 | 77 | 358 | 117 | 406 | UDP-glucuronosyltransferase 2B13 OS=Oryctolagus cuniculus OX=9986 GN=UGT2B13 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
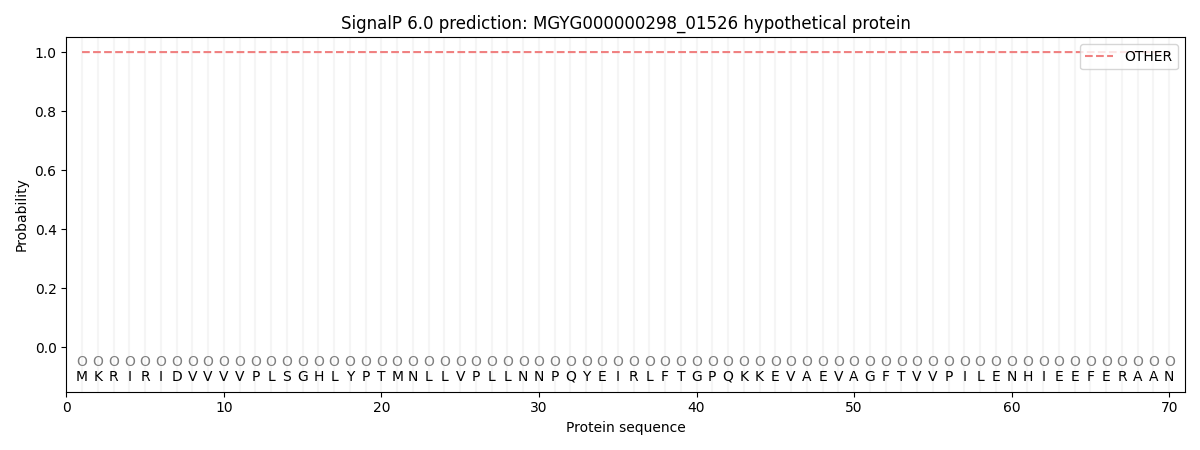
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000072 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
