You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000299_01389
You are here: Home > Sequence: MGYG000000299_01389
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pauljensenia sp900541895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Pauljensenia; Pauljensenia sp900541895 | |||||||||||
| CAZyme ID | MGYG000000299_01389 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9045; End: 9950 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT1 | 39 | 294 | 6.6e-25 | 0.6832460732984293 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1819 | YjiC | 7.34e-23 | 1 | 298 | 67 | 403 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| cd03784 | GT1_Gtf-like | 3.20e-16 | 17 | 293 | 89 | 404 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| TIGR00661 | MJ1255 | 1.75e-07 | 14 | 253 | 83 | 305 | conserved hypothetical protein. This model represents nearly the full length of MJ1255 from Methanococcus jannaschii and of an unpublished protein from Vibrio cholerae, as well as the C-terminal half of a protein from Methanobacterium thermoautotrophicum. A small region (~50 amino acids) within the domain appears related to a family of sugar transferases. [Hypothetical proteins, Conserved] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCT36430.1 | 2.84e-163 | 1 | 294 | 85 | 376 |
| QGS11935.1 | 7.30e-159 | 1 | 294 | 85 | 376 |
| AHF25143.1 | 7.79e-79 | 1 | 291 | 105 | 402 |
| BAR06867.1 | 4.45e-63 | 1 | 294 | 93 | 383 |
| BAR05900.1 | 5.20e-60 | 1 | 295 | 121 | 417 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3OTG_A | 2.56e-17 | 16 | 288 | 118 | 401 | CrystalStructure of CalG1, Calicheamicin Glycostyltransferase, TDP bound form [Micromonospora echinospora],3OTH_A Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora],3OTH_B Crystal Structure of CalG1, Calicheamicin Glycostyltransferase, TDP and calicheamicin alpha3I bound form [Micromonospora echinospora] |
| 4AMB_A | 1.22e-12 | 11 | 280 | 111 | 385 | Crystalstructure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AMB_B Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater] |
| 4AMG_A | 1.22e-12 | 11 | 280 | 111 | 385 | Crystalstructure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AMG_B Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AN4_A Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AN4_B Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AN4_C Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater],4AN4_D Crystal structure of the glycosyltransferase SnogD from Streptomyces nogalater [Streptomyces nogalater] |
| 4RIE_A | 7.82e-09 | 17 | 295 | 96 | 374 | LandomycinGlycosyltransferase LanGT2 [Streptomyces cyanogenus],4RIE_B Landomycin Glycosyltransferase LanGT2 [Streptomyces cyanogenus],4RIF_A Landomycin Glycosyltransferase LanGT2, carbasugar substrate complex [Streptomyces cyanogenus],4RIF_B Landomycin Glycosyltransferase LanGT2, carbasugar substrate complex [Streptomyces cyanogenus] |
| 6KQW_A | 7.96e-09 | 192 | 294 | 282 | 385 | ChainA, Uncharacterized UDP-glucosyltransferase YjiC [Bacillus subtilis subsp. subtilis str. 168] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9F2F9 | 1.64e-10 | 4 | 280 | 87 | 359 | Elloramycin glycosyltransferase ElmGT OS=Streptomyces olivaceus OX=47716 GN=elmGT PE=1 SV=2 |
| Q9L555 | 2.01e-08 | 83 | 293 | 217 | 422 | Aclacinomycin-T 2-deoxy-L-fucose transferase OS=Streptomyces galilaeus OX=33899 GN=aknK PE=1 SV=1 |
| O34539 | 4.41e-08 | 192 | 294 | 282 | 385 | NDP-glycosyltransferase YjiC OS=Bacillus subtilis (strain 168) OX=224308 GN=yjiC PE=1 SV=1 |
| Q53685 | 2.04e-06 | 133 | 301 | 251 | 410 | Oleandomycin glycosyltransferase OS=Streptomyces antibioticus OX=1890 GN=oleD PE=1 SV=1 |
| Q9L9F5 | 2.50e-06 | 20 | 245 | 94 | 322 | L-demethylnoviosyl transferase OS=Streptomyces niveus OX=193462 GN=novM PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
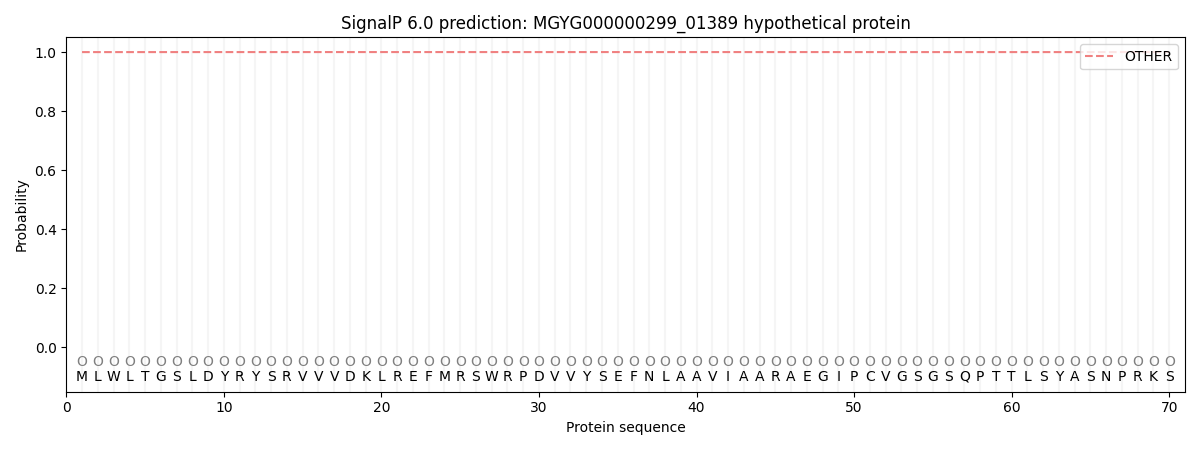
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000069 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
