You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000311_01345
You are here: Home > Sequence: MGYG000000311_01345
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Veillonella sp900550175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Veillonellales; Veillonellaceae; Veillonella; Veillonella sp900550175 | |||||||||||
| CAZyme ID | MGYG000000311_01345 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39; End: 1520 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 4 | 468 | 4.1e-71 | 0.8685185185185185 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1807 | ArnT | 8.62e-30 | 1 | 368 | 55 | 422 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| PRK13279 | arnT | 1.84e-10 | 7 | 322 | 59 | 364 | lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
| pfam13231 | PMT_2 | 6.03e-09 | 9 | 127 | 1 | 118 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQB17357.1 | 1.04e-310 | 1 | 493 | 54 | 546 |
| SNV00837.1 | 1.04e-310 | 1 | 493 | 54 | 546 |
| ACZ25181.1 | 1.04e-310 | 1 | 493 | 54 | 546 |
| CAB1276937.1 | 1.16e-307 | 1 | 493 | 54 | 546 |
| VEG94157.1 | 3.61e-222 | 1 | 493 | 54 | 538 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EZM_A | 1.30e-11 | 7 | 289 | 85 | 367 | CrystalStructure of ArnT from Cupriavidus metallidurans in the apo state [Cupriavidus metallidurans CH34],5F15_A Crystal Structure of ArnT from Cupriavidus metallidurans bound to Undecaprenyl phosphate [Cupriavidus metallidurans CH34] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O67270 | 1.27e-19 | 9 | 274 | 58 | 309 | Uncharacterized protein aq_1220 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1220 PE=3 SV=1 |
| O67601 | 1.12e-17 | 8 | 275 | 57 | 299 | Uncharacterized protein aq_1704 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1704 PE=3 SV=1 |
| B4EUL1 | 1.07e-14 | 4 | 289 | 59 | 336 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 1 OS=Proteus mirabilis (strain HI4320) OX=529507 GN=arnT1 PE=3 SV=1 |
| Q4KC80 | 5.89e-14 | 7 | 315 | 58 | 355 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 1 OS=Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5) OX=220664 GN=arnT1 PE=3 SV=1 |
| Q3KCB9 | 2.92e-11 | 7 | 315 | 58 | 355 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 2 OS=Pseudomonas fluorescens (strain Pf0-1) OX=205922 GN=arnT2 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
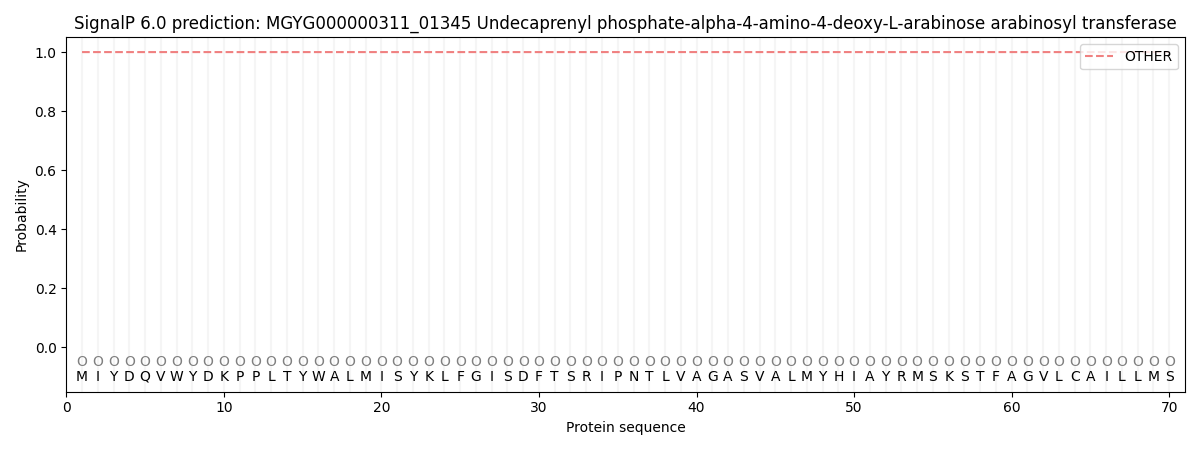
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
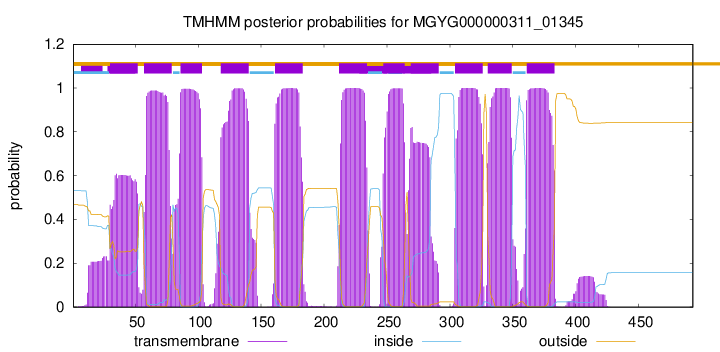
| start | end |
|---|---|
| 30 | 52 |
| 57 | 79 |
| 86 | 103 |
| 118 | 140 |
| 161 | 183 |
| 212 | 234 |
| 247 | 264 |
| 269 | 291 |
| 304 | 326 |
| 330 | 349 |
| 361 | 383 |
