You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000323_00424
You are here: Home > Sequence: MGYG000000323_00424
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
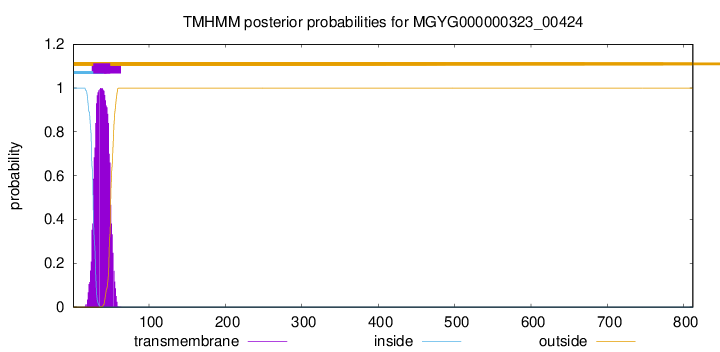
TMHMM annotations
Basic Information help
| Species | Enterococcus_D sp002850555 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus_D; Enterococcus_D sp002850555 | |||||||||||
| CAZyme ID | MGYG000000323_00424 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Monofunctional biosynthetic peptidoglycan transglycosylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 443348; End: 445786 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 86 | 273 | 1.6e-52 | 0.9887005649717514 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0744 | MrcB | 5.42e-150 | 15 | 684 | 4 | 600 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| COG5009 | MrcA | 4.94e-91 | 25 | 737 | 2 | 785 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
| pfam00912 | Transgly | 1.45e-55 | 86 | 274 | 1 | 177 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
| COG4953 | PbpC | 6.03e-43 | 49 | 684 | 8 | 557 | Membrane carboxypeptidase/penicillin-binding protein PbpC [Cell wall/membrane/envelope biogenesis]. |
| PRK14850 | PRK14850 | 7.95e-38 | 99 | 665 | 157 | 673 | penicillin-binding protein 1b; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATF70746.1 | 0.0 | 1 | 773 | 1 | 773 |
| AMG50991.1 | 0.0 | 1 | 772 | 1 | 772 |
| AUJ85904.1 | 0.0 | 1 | 772 | 1 | 772 |
| EEV38408.2 | 0.0 | 1 | 773 | 1 | 773 |
| QGN29249.1 | 0.0 | 1 | 772 | 1 | 772 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2JE5_A | 1.42e-219 | 48 | 770 | 3 | 718 | StructuralAnd Mechanistic Basis Of Penicillin Binding Protein Inhibition By Lactivicins [Streptococcus pneumoniae R6],2JE5_B Structural And Mechanistic Basis Of Penicillin Binding Protein Inhibition By Lactivicins [Streptococcus pneumoniae R6] |
| 2XD1_A | 1.21e-117 | 292 | 770 | 24 | 492 | ACTIVESITE RESTRUCTURING REGULATES LIGAND RECOGNITION IN CLASS A PENICILLIN-BINDING PROTEINS [Streptococcus pneumoniae R6],2XD1_B ACTIVE SITE RESTRUCTURING REGULATES LIGAND RECOGNITION IN CLASS A PENICILLIN-BINDING PROTEINS [Streptococcus pneumoniae R6] |
| 2BG1_A | 1.21e-117 | 292 | 770 | 24 | 492 | Activesite restructuring regulates ligand recognition in classA Penicillin-binding proteins (PBPs) [Streptococcus pneumoniae R6],2XD5_A Structural insights into the catalytic mechanism and the role of Streptococcus pneumoniae PBP1b [Streptococcus pneumoniae R6],2XD5_B Structural insights into the catalytic mechanism and the role of Streptococcus pneumoniae PBP1b [Streptococcus pneumoniae R6] |
| 2UWX_A | 6.65e-117 | 292 | 770 | 24 | 492 | Activesite restructuring regulates ligand recognition in class A penicillin-binding proteins [Streptococcus pneumoniae R6] |
| 2Y2G_A | 3.65e-116 | 292 | 770 | 24 | 492 | Penicillin-BindingProtein 1b (Pbp-1b) In Complex With An Alkyl Boronate (A01) [Streptococcus pneumoniae R6],2Y2G_B Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (A01) [Streptococcus pneumoniae R6],2Y2H_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Za2) [Streptococcus pneumoniae R6],2Y2H_B Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Za2) [Streptococcus pneumoniae R6],2Y2I_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Za3) [Streptococcus pneumoniae R6],2Y2J_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Za4) [Streptococcus pneumoniae R6],2Y2K_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Za5) [Streptococcus pneumoniae R6],2Y2L_A Penicillin-binding Protein 1b (pbp-1b) In Complex With An Alkyl Boronate (e06) [Streptococcus pneumoniae R6],2Y2L_B Penicillin-binding Protein 1b (pbp-1b) In Complex With An Alkyl Boronate (e06) [Streptococcus pneumoniae R6],2Y2M_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (E08) [Streptococcus pneumoniae R6],2Y2N_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (E07) [Streptococcus pneumoniae R6],2Y2O_A Penicillin-binding Protein 1b (pbp-1b) In Complex With An Alkyl Boronate (eo9) [Streptococcus pneumoniae R6],2Y2P_A Penicillin-binding protein 1b (pbp-1b) in complex with an alkyl boronate (z10) [Streptococcus pneumoniae R6],2Y2Q_A Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Z06) [Streptococcus pneumoniae R6],2Y2Q_B Penicillin-Binding Protein 1b (Pbp-1b) In Complex With An Alkyl Boronate (Z06) [Streptococcus pneumoniae R6] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39793 | 2.29e-57 | 47 | 664 | 47 | 619 | Penicillin-binding protein 1A/1B OS=Bacillus subtilis (strain 168) OX=224308 GN=ponA PE=1 SV=1 |
| Q0SRL7 | 7.92e-55 | 43 | 726 | 34 | 702 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain SM101 / Type A) OX=289380 GN=pbpA PE=3 SV=1 |
| Q0TNZ8 | 1.81e-53 | 4 | 726 | 9 | 702 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=pbpA PE=3 SV=1 |
| Q8XJ01 | 1.97e-53 | 43 | 674 | 34 | 655 | Penicillin-binding protein 1A OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=pbpA PE=3 SV=1 |
| Q891X1 | 1.03e-52 | 84 | 718 | 64 | 707 | Penicillin-binding protein 1A OS=Clostridium tetani (strain Massachusetts / E88) OX=212717 GN=pbpA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
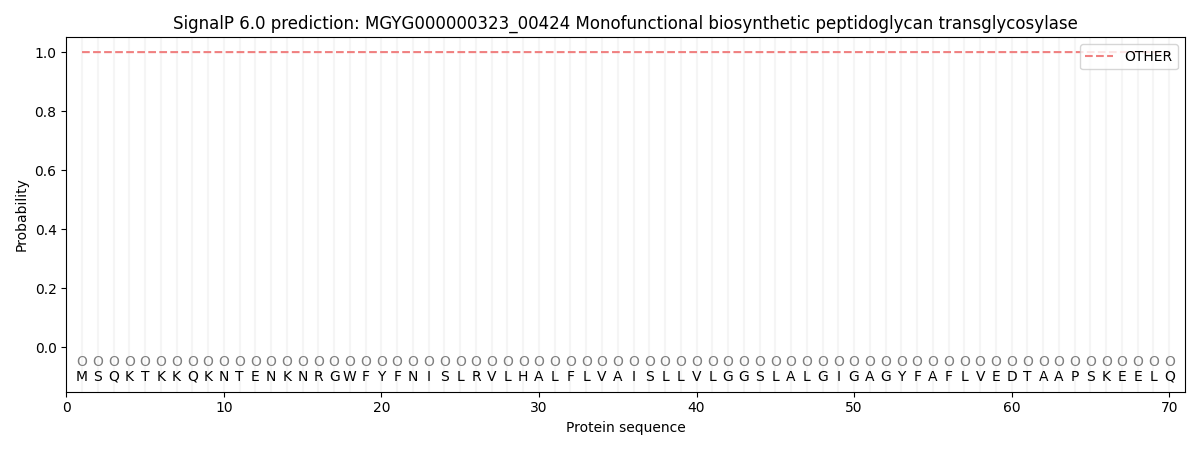
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999995 | 0.000017 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |