You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000339_00039
You are here: Home > Sequence: MGYG000000339_00039
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
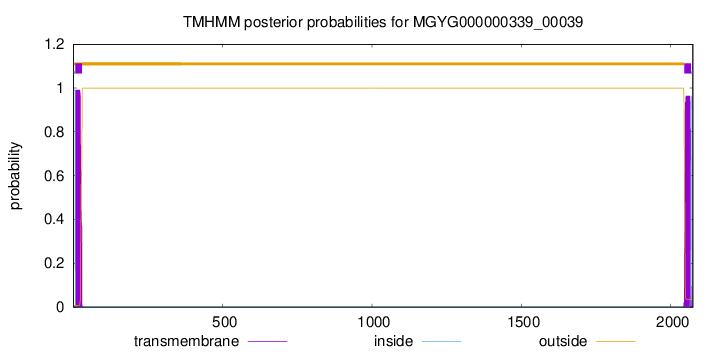
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; UBA9475; | |||||||||||
| CAZyme ID | MGYG000000339_00039 | |||||||||||
| CAZy Family | GH110 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5718; End: 11942 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH110 | 41 | 636 | 9.8e-140 | 0.958029197080292 |
| CBM51 | 1582 | 1716 | 3.9e-36 | 0.9328358208955224 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08305 | NPCBM | 3.32e-38 | 1573 | 1717 | 4 | 136 | NPCBM/NEW2 domain. This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. This domain has also been called the NEW2 domain (Naumoff DG. Phylogenetic analysis of alpha-galactosidases of the GH27 family. Molecular Biology (Engl Transl). (2004)38:388-399.) |
| smart00776 | NPCBM | 3.87e-28 | 1568 | 1716 | 1 | 144 | This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. |
| COG5492 | YjdB | 7.81e-21 | 1725 | 1822 | 178 | 279 | Uncharacterized conserved protein YjdB, contains Ig-like domain [General function prediction only]. |
| pfam02368 | Big_2 | 1.33e-15 | 1729 | 1805 | 1 | 77 | Bacterial Ig-like domain (group 2). This family consists of bacterial domains with an Ig-like fold. Members of this family are found in bacterial and phage surface proteins such as intimins. |
| smart00635 | BID_2 | 1.04e-10 | 1727 | 1804 | 1 | 81 | Bacterial Ig-like domain 2. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AYE35188.1 | 5.41e-249 | 24 | 1399 | 25 | 1395 |
| QAS60591.1 | 2.06e-248 | 24 | 1399 | 25 | 1395 |
| ATD55583.1 | 4.65e-169 | 42 | 714 | 3 | 618 |
| ATD56740.1 | 4.65e-169 | 42 | 714 | 3 | 618 |
| QBJ75880.1 | 4.97e-169 | 42 | 714 | 5 | 620 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7JW4_A | 1.57e-46 | 19 | 606 | 12 | 542 | Crystalstructure of PdGH110B in complex with D-galactose [Pseudoalteromonas distincta],7JW4_B Crystal structure of PdGH110B in complex with D-galactose [Pseudoalteromonas distincta] |
| 7JWF_A | 6.86e-46 | 19 | 606 | 12 | 542 | Crystalstructure of PdGH110B D344N in complex with alpha-(1,3)-galactobiose [Pseudoalteromonas distincta],7JWF_B Crystal structure of PdGH110B D344N in complex with alpha-(1,3)-galactobiose [Pseudoalteromonas distincta],7JWF_C Crystal structure of PdGH110B D344N in complex with alpha-(1,3)-galactobiose [Pseudoalteromonas distincta],7JWF_D Crystal structure of PdGH110B D344N in complex with alpha-(1,3)-galactobiose [Pseudoalteromonas distincta] |
| 7JS4_A | 7.68e-27 | 1561 | 1811 | 600 | 833 | ChainA, F5/8 type C domain protein [Clostridium perfringens ATCC 13124] |
| 7JRM_A | 2.66e-24 | 1543 | 1716 | 55 | 223 | ChainA, F5/8 type C domain protein [Clostridium perfringens ATCC 13124] |
| 2VMH_A | 2.95e-24 | 1587 | 1715 | 22 | 149 | Thestructure of CBM51 from Clostridium perfringens GH95 [Clostridium perfringens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B1V8K7 | 1.17e-118 | 40 | 774 | 33 | 723 | Alpha-1,3-galactosidase A OS=Streptacidiphilus griseoplanus OX=66896 GN=glaA PE=1 SV=1 |
| Q826C5 | 4.08e-101 | 6 | 693 | 12 | 624 | Alpha-1,3-galactosidase A OS=Streptomyces avermitilis (strain ATCC 31267 / DSM 46492 / JCM 5070 / NBRC 14893 / NCIMB 12804 / NRRL 8165 / MA-4680) OX=227882 GN=glaA PE=3 SV=1 |
| Q8A2Z5 | 3.35e-60 | 59 | 629 | 29 | 532 | Alpha-1,3-galactosidase A OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=glaA PE=3 SV=1 |
| Q89ZX0 | 2.96e-59 | 37 | 632 | 23 | 556 | Alpha-1,3-galactosidase B OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=glaB PE=3 SV=1 |
| Q64MU6 | 3.59e-58 | 42 | 631 | 26 | 552 | Alpha-1,3-galactosidase A OS=Bacteroides fragilis (strain YCH46) OX=295405 GN=glaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
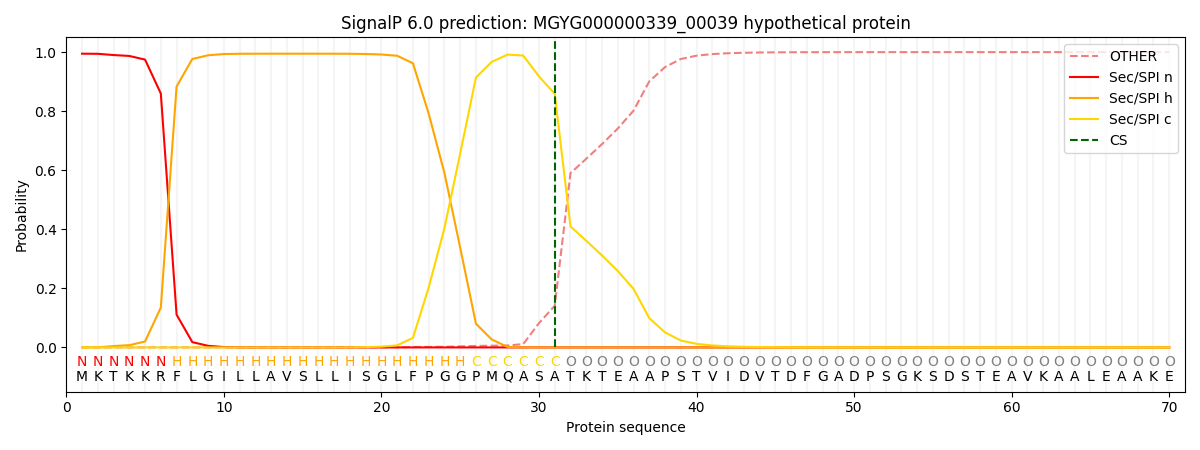
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000586 | 0.992756 | 0.005881 | 0.000348 | 0.000210 | 0.000167 |