You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000339_00092
You are here: Home > Sequence: MGYG000000339_00092
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
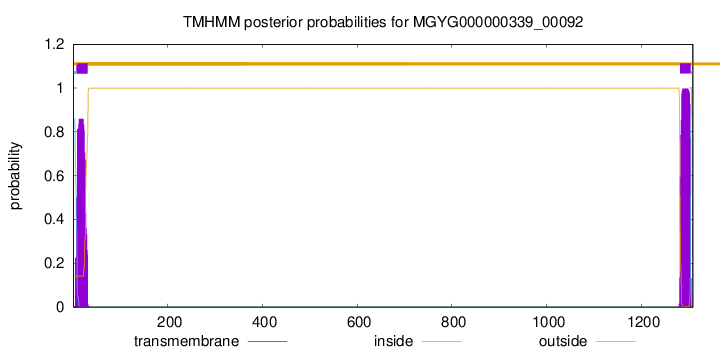
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; UBA9475; | |||||||||||
| CAZyme ID | MGYG000000339_00092 | |||||||||||
| CAZy Family | GH51 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8939; End: 12868 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3534 | AbfA | 3.26e-26 | 370 | 956 | 66 | 498 | Alpha-L-arabinofuranosidase [Carbohydrate transport and metabolism]. |
| smart00813 | Alpha-L-AF_C | 1.63e-16 | 831 | 950 | 67 | 189 | Alpha-L-arabinofuranosidase C-terminus. This entry represents the C terminus (approximately 200 residues) of bacterial and eukaryotic alpha-L-arabinofuranosidase. This catalyses the hydrolysis of non-reducing terminal alpha-L-arabinofuranosidic linkages in L-arabinose-containing polysaccharides. |
| pfam18998 | Flg_new_2 | 7.96e-14 | 964 | 1033 | 1 | 74 | Divergent InlB B-repeat domain. This family of domains are found in bacterial cell surface proteins. They are often found in tandem array. This domain is closely related to pfam09479. |
| pfam13385 | Laminin_G_3 | 1.40e-13 | 82 | 235 | 2 | 149 | Concanavalin A-like lectin/glucanases superfamily. This domain belongs to the Concanavalin A-like lectin/glucanases superfamily. |
| pfam06964 | Alpha-L-AF_C | 2.24e-13 | 831 | 950 | 70 | 192 | Alpha-L-arabinofuranosidase C-terminal domain. This family represents the C-terminus (approximately 200 residues) of bacterial and eukaryotic alpha-L-arabinofuranosidase (EC:3.2.1.55). This catalyzes the hydrolysis of nonreducing terminal alpha-L-arabinofuranosidic linkages in L-arabinose-containing polysaccharides. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADO52227.1 | 0.0 | 30 | 963 | 45 | 996 |
| BBA56144.1 | 0.0 | 30 | 963 | 28 | 979 |
| BAQ97226.1 | 0.0 | 30 | 963 | 28 | 979 |
| VEG16797.1 | 0.0 | 30 | 963 | 45 | 996 |
| QRI58751.1 | 0.0 | 30 | 963 | 45 | 996 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5O7Z_A | 4.18e-12 | 345 | 488 | 40 | 173 | ChainA, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O7Z_B Chain B, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O7Z_C Chain C, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O7Z_D Chain D, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O7Z_E Chain E, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O7Z_F Chain F, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_A Chain A, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_B Chain B, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_C Chain C, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_D Chain D, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_E Chain E, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila],5O80_F Chain F, Intracellular exo-alpha-(1->5)-L-arabinofuranosidase [Thermochaetoides thermophila] |
| 2C7F_A | 4.32e-12 | 345 | 488 | 51 | 184 | ChainA, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C7F_B Chain B, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C7F_C Chain C, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C7F_D Chain D, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C7F_E Chain E, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C7F_F Chain F, Alpha-l-arabinofuranosidase [Acetivibrio thermocellus],2C8N_A Chain A, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus],2C8N_B Chain B, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus],2C8N_C Chain C, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus],2C8N_D Chain D, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus],2C8N_E Chain E, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus],2C8N_F Chain F, ALPHA-L-ARABINOFURANOSIDASE [Acetivibrio thermocellus] |
| 4ATW_A | 6.88e-12 | 369 | 488 | 64 | 173 | Thecrystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_B The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_C The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_D The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_E The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8],4ATW_F The crystal structure of Arabinofuranosidase [Thermotoga maritima MSB8] |
| 3S2C_A | 6.92e-12 | 369 | 488 | 64 | 173 | Structureof the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_B Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_C Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_D Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_E Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_F Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_G Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_H Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_I Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_J Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_K Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3S2C_L Structure of the thermostable GH51 alpha-L-arabinofuranosidase from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1] |
| 3UG3_A | 7.34e-12 | 369 | 488 | 84 | 193 | Crystalstructure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG3_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima ligand free form [Thermotoga maritima],3UG4_A Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG4_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima arabinose complex [Thermotoga maritima],3UG5_A Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_B Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_C Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_D Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_E Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima],3UG5_F Crystal structure of alpha-L-arabinofuranosidase from Thermotoga maritima xylose complex [Thermotoga maritima] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DIH0 | 4.34e-12 | 345 | 488 | 41 | 174 | Intracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=Cthe_2548 PE=1 SV=1 |
| P53627 | 6.30e-10 | 343 | 488 | 41 | 176 | Intracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Streptomyces lividans OX=1916 GN=abfA PE=1 SV=1 |
| P94531 | 1.46e-09 | 370 | 488 | 66 | 174 | Intracellular exo-alpha-(1->5)-L-arabinofuranosidase 1 OS=Bacillus subtilis (strain 168) OX=224308 GN=abfA PE=1 SV=2 |
| Q841V6 | 2.88e-09 | 355 | 488 | 93 | 216 | Intracellular exo-alpha-(1->5)-L-arabinofuranosidase OS=Bifidobacterium longum OX=216816 GN=abfB PE=1 SV=1 |
| B8NIX4 | 4.46e-09 | 370 | 487 | 75 | 182 | Probable alpha-L-arabinofuranosidase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=abfC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.003889 | 0.510668 | 0.484460 | 0.000400 | 0.000280 | 0.000280 |