You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000339_00657
You are here: Home > Sequence: MGYG000000339_00657
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; UBA9475; | |||||||||||
| CAZyme ID | MGYG000000339_00657 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 556; End: 3483 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 417 | 841 | 8.5e-57 | 0.4441489361702128 |
| GH20 | 6 | 209 | 1.4e-25 | 0.5905044510385756 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06564 | GH20_DspB_LnbB-like | 6.16e-74 | 1 | 209 | 129 | 325 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| COG3250 | LacZ | 9.40e-33 | 441 | 706 | 156 | 408 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10340 | ebgA | 2.08e-30 | 449 | 707 | 195 | 452 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK09525 | lacZ | 1.64e-21 | 450 | 660 | 208 | 412 | beta-galactosidase. |
| PRK10150 | PRK10150 | 1.70e-18 | 448 | 704 | 167 | 422 | beta-D-glucuronidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPS14026.1 | 5.56e-116 | 2 | 427 | 703 | 1113 |
| QQV05894.1 | 5.56e-116 | 2 | 427 | 703 | 1113 |
| QQY27155.1 | 5.56e-116 | 2 | 427 | 703 | 1113 |
| QMW75640.1 | 5.56e-116 | 2 | 427 | 703 | 1113 |
| AYE35153.1 | 5.68e-97 | 1 | 440 | 562 | 984 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6JQF_A | 6.65e-48 | 12 | 411 | 355 | 721 | Crystallizationanalysis of a beta-N-acetylhexosaminidase (Am2136) from Akkermansia muciniphila [Akkermansia muciniphila ATCC BAA-835] |
| 4YPJ_A | 3.52e-29 | 326 | 928 | 61 | 658 | ChainA, Beta galactosidase [Niallia circulans],4YPJ_B Chain B, Beta galactosidase [Niallia circulans] |
| 6HPD_A | 4.82e-29 | 355 | 912 | 112 | 650 | Thestructure of a beta-glucuronidase from glycoside hydrolase family 2 [Formosa agariphila KMM 3901] |
| 7CWD_A | 5.72e-28 | 326 | 928 | 55 | 652 | ChainA, beta-glalactosidase [Niallia circulans],7CWI_A Chain A, beta-galactosidase [Niallia circulans] |
| 5T98_A | 1.22e-26 | 376 | 915 | 103 | 650 | Crystalstructure of BuGH2Awt [Bacteroides uniformis],5T98_B Crystal structure of BuGH2Awt [Bacteroides uniformis],5T99_A Crystal structure of BuGH2Awt in complex with Galactoisofagomine [Bacteroides uniformis],5T99_B Crystal structure of BuGH2Awt in complex with Galactoisofagomine [Bacteroides uniformis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B2UPR7 | 2.31e-49 | 12 | 411 | 377 | 743 | Beta-hexosaminidase Amuc_2136 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_2136 PE=1 SV=1 |
| P77989 | 6.99e-35 | 448 | 917 | 146 | 586 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
| T2KPJ7 | 9.39e-30 | 368 | 926 | 132 | 686 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| T2KN75 | 2.61e-28 | 355 | 912 | 103 | 641 | Beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22060 PE=1 SV=1 |
| P26257 | 3.86e-26 | 448 | 905 | 144 | 583 | Beta-galactosidase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=lacZ PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
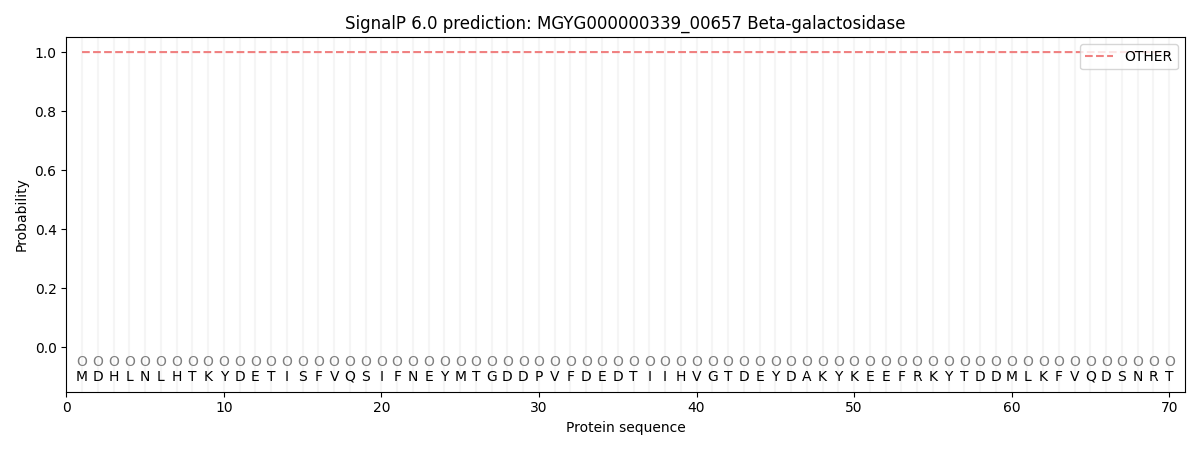
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000075 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
