You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000345_00478
You are here: Home > Sequence: MGYG000000345_00478
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1829 sp002338895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; UBA1829; UBA1829 sp002338895 | |||||||||||
| CAZyme ID | MGYG000000345_00478 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 228593; End: 229909 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 77 | 377 | 5.7e-46 | 0.9141914191419142 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 1.08e-40 | 123 | 375 | 17 | 260 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.28e-33 | 77 | 375 | 22 | 305 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 3.80e-29 | 99 | 384 | 69 | 344 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM44415.1 | 3.19e-204 | 1 | 429 | 3 | 431 |
| QHV94149.1 | 2.61e-105 | 16 | 438 | 28 | 445 |
| QIP12711.1 | 3.69e-105 | 16 | 438 | 28 | 445 |
| QMW04637.1 | 4.32e-101 | 16 | 438 | 28 | 446 |
| AEI51978.1 | 4.31e-100 | 15 | 437 | 27 | 440 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NIY_A | 4.02e-23 | 71 | 383 | 39 | 337 | Crystalstructure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NIY_B Crystal structure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NJ3_A Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1],3NJ3_B Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1] |
| 1VBR_A | 4.53e-23 | 71 | 383 | 23 | 321 | Crystalstructure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBR_B Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBU_A Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima],1VBU_B Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima] |
| 7D88_A | 7.31e-21 | 46 | 436 | 71 | 421 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
| 7NL2_A | 1.59e-20 | 85 | 383 | 30 | 343 | ChainA, Beta-xylanase [Pseudothermotoga thermarum DSM 5069],7NL2_B Chain B, Beta-xylanase [Pseudothermotoga thermarum DSM 5069] |
| 7D89_A | 5.90e-20 | 46 | 436 | 71 | 421 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q12603 | 4.10e-21 | 82 | 325 | 57 | 287 | Beta-1,4-xylanase OS=Dictyoglomus thermophilum OX=14 GN=xynA PE=3 SV=1 |
| Q60041 | 8.12e-20 | 119 | 383 | 82 | 340 | Endo-1,4-beta-xylanase B OS=Thermotoga neapolitana OX=2337 GN=xynB PE=3 SV=1 |
| O80596 | 2.15e-18 | 43 | 428 | 706 | 1052 | Endo-1,4-beta-xylanase 2 OS=Arabidopsis thaliana OX=3702 GN=XYN2 PE=3 SV=1 |
| Q02290 | 6.16e-18 | 118 | 383 | 74 | 326 | Endo-1,4-beta-xylanase B OS=Neocallimastix patriciarum OX=4758 GN=xynB PE=2 SV=1 |
| P40944 | 7.09e-18 | 125 | 326 | 415 | 625 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor sp. (strain Rt8B.4) OX=28238 GN=xynA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
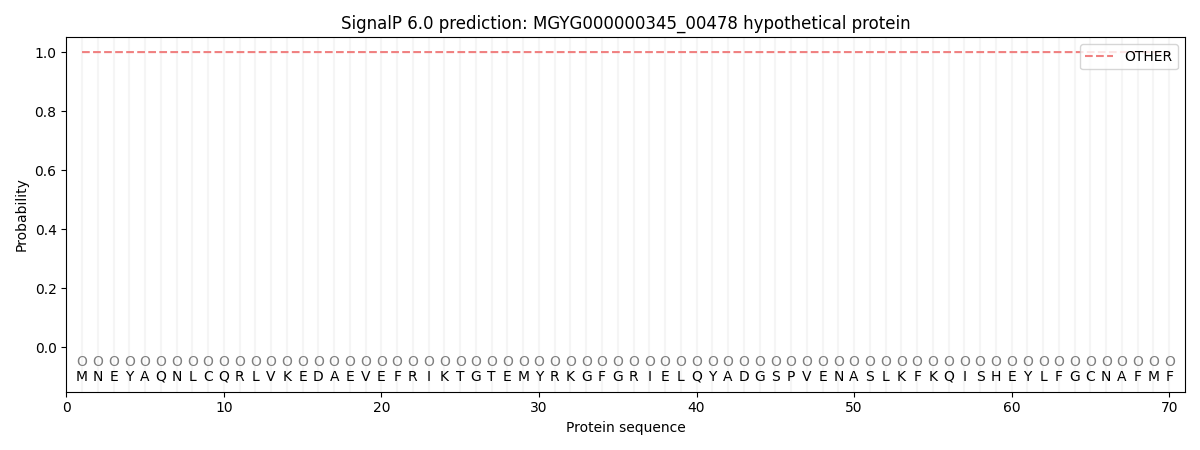
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000047 | 0.000003 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
