You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000345_02899
You are here: Home > Sequence: MGYG000000345_02899
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1829 sp002338895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; UBA1829; UBA1829 sp002338895 | |||||||||||
| CAZyme ID | MGYG000000345_02899 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-glucuronidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11311; End: 12990 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 15 | 556 | 1.7e-78 | 0.6050531914893617 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10150 | PRK10150 | 2.01e-62 | 15 | 553 | 15 | 590 | beta-D-glucuronidase; Provisional |
| COG3250 | LacZ | 1.88e-53 | 15 | 556 | 15 | 596 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| pfam02836 | Glyco_hydro_2_C | 5.95e-35 | 251 | 556 | 1 | 298 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK10340 | ebgA | 6.22e-28 | 140 | 421 | 190 | 487 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK09525 | lacZ | 4.63e-23 | 146 | 375 | 208 | 465 | beta-galactosidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM46586.1 | 3.73e-154 | 1 | 555 | 1 | 558 |
| AVM47139.1 | 1.75e-138 | 1 | 555 | 64 | 622 |
| AHF91981.1 | 1.69e-128 | 6 | 555 | 22 | 589 |
| QZD54105.1 | 7.10e-124 | 2 | 555 | 3 | 553 |
| QYY34828.1 | 4.04e-123 | 15 | 555 | 11 | 560 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6NCW_A | 2.64e-95 | 45 | 555 | 39 | 552 | Crystalstructure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_B Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_C Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCW_D Crystal structure of a GH2 beta-galacturonidase from Eisenbergiella tayi bound to glycerol [Eisenbergiella tayi],6NCX_A Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_B Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_C Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi],6NCX_D Crystal structure of GH2 beta-galacturonidase from Eisenbergiella tayi bound to galacturonate [Eisenbergiella tayi] |
| 6U7I_A | 1.09e-49 | 130 | 556 | 170 | 589 | Faecalibacteriumprausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_B Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_C Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_D Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 6BO6_A | 3.48e-49 | 12 | 556 | 12 | 601 | Eubacteriumeligens beta-glucuronidase bound to UNC4917 glucuronic acid conjugate [[Eubacterium] eligens ATCC 27750] |
| 6BJQ_A | 4.77e-49 | 12 | 556 | 36 | 625 | ChainA, Glycoside Hydrolase Family 2 candidate b-glucuronidase [[Eubacterium] eligens ATCC 27750] |
| 6BJW_A | 4.83e-49 | 12 | 556 | 36 | 625 | ChainA, Glycoside Hydrolase Family 2 candidate b-glucuronidase [[Eubacterium] eligens ATCC 27750] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P05804 | 3.59e-42 | 15 | 556 | 15 | 589 | Beta-glucuronidase OS=Escherichia coli (strain K12) OX=83333 GN=uidA PE=1 SV=2 |
| P06760 | 1.50e-37 | 139 | 555 | 205 | 622 | Beta-glucuronidase OS=Rattus norvegicus OX=10116 GN=Gusb PE=1 SV=1 |
| P12265 | 2.03e-37 | 139 | 555 | 205 | 622 | Beta-glucuronidase OS=Mus musculus OX=10090 GN=Gusb PE=1 SV=2 |
| T2KPJ7 | 2.19e-35 | 47 | 556 | 89 | 629 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| O18835 | 2.18e-34 | 128 | 555 | 190 | 625 | Beta-glucuronidase OS=Canis lupus familiaris OX=9615 GN=GUSB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
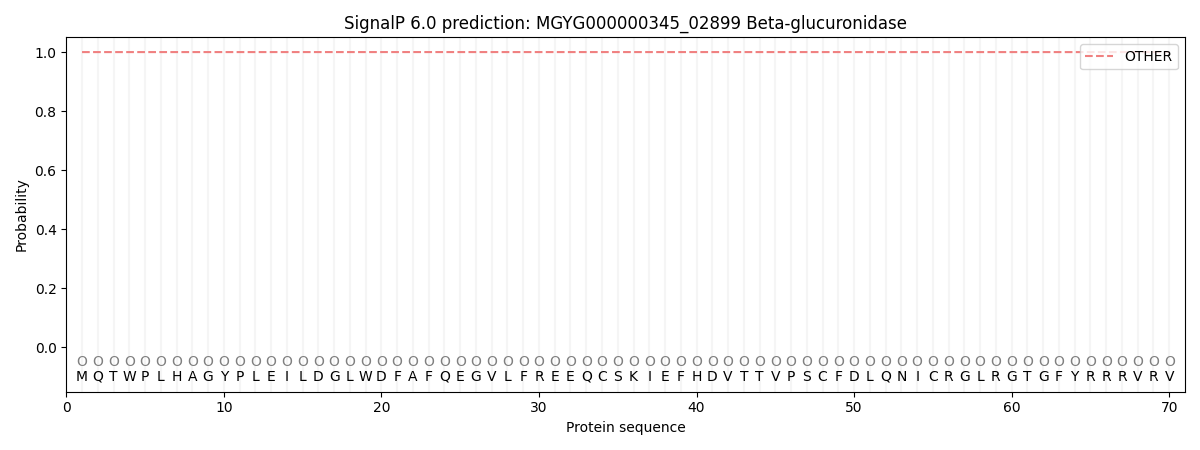
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000079 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
