You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000355_00035
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides;
|
| CAZyme ID |
MGYG000000355_00035
|
| CAZy Family |
GT83 |
| CAZyme Description |
Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 275 |
|
30933.55 |
7.9288 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000355 |
2806154 |
MAG |
Sweden |
Europe |
|
| Gene Location |
Start: 27052;
End: 27879
Strand: +
|
No EC number prediction in MGYG000000355_00035.
| Family |
Start |
End |
Evalue |
family coverage |
| GT83 |
3 |
262 |
3.1e-51 |
0.4666666666666667 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| COG1807
|
ArnT |
2.36e-23 |
1 |
260 |
4 |
262 |
4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| PRK13279
|
arnT |
1.20e-04 |
1 |
224 |
2 |
224 |
lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
| COG4745
|
COG4745 |
0.005 |
35 |
175 |
42 |
177 |
Predicted membrane-bound mannosyltransferase [General function prediction only]. |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
|
O67270
|
2.47e-11 |
11 |
260 |
6 |
250 |
Uncharacterized protein aq_1220 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1220 PE=3 SV=1 |
|
O67601
|
1.29e-10 |
1 |
260 |
1 |
240 |
Uncharacterized protein aq_1704 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1704 PE=3 SV=1 |
This protein is predicted as OTHER

| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.999981
|
0.000034
|
0.000008
|
0.000000
|
0.000000
|
0.000003
|
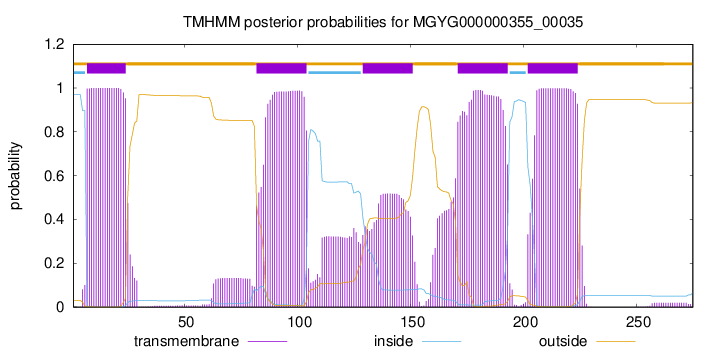
| start |
end |
| 7 |
24 |
| 82 |
104 |
| 129 |
151 |
| 171 |
193 |
| 202 |
224 |


