You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000362_01437
You are here: Home > Sequence: MGYG000000362_01437
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-115 sp003531585 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; CAG-115; CAG-115 sp003531585 | |||||||||||
| CAZyme ID | MGYG000000362_01437 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 22600; End: 24201 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 69 | 260 | 1.2e-38 | 0.7159533073929961 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 2.08e-13 | 38 | 285 | 1 | 223 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.31e-12 | 70 | 180 | 76 | 191 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| COG3934 | COG3934 | 1.24e-04 | 71 | 462 | 30 | 384 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AQG79101.1 | 1.46e-117 | 33 | 529 | 45 | 550 |
| ABG59109.1 | 6.01e-88 | 31 | 529 | 36 | 549 |
| QSI77041.1 | 4.93e-60 | 26 | 267 | 40 | 267 |
| ACR11279.1 | 1.13e-52 | 28 | 265 | 28 | 251 |
| ASM78055.1 | 2.91e-44 | 24 | 512 | 183 | 707 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1CEC_A | 1.43e-30 | 57 | 267 | 13 | 210 | ChainA, ENDOGLUCANASE CELC [Acetivibrio thermocellus] |
| 1CEN_A | 3.66e-30 | 57 | 267 | 13 | 210 | ChainA, CELLULASE CELC [Acetivibrio thermocellus],1CEO_A Chain A, CELLULASE CELC [Acetivibrio thermocellus] |
| 7EC9_A | 9.67e-18 | 45 | 286 | 16 | 245 | ChainA, Endoglucanase [Thermotoga maritima MSB8],7EC9_B Chain B, Endoglucanase [Thermotoga maritima MSB8],7EFZ_A Chain A, Endoglucanase [Thermotoga maritima MSB8],7EFZ_B Chain B, Endoglucanase [Thermotoga maritima MSB8] |
| 1VJZ_A | 2.49e-16 | 45 | 286 | 16 | 245 | Crystalstructure of Endoglucanase (TM1752) from Thermotoga maritima at 2.05 A resolution [Thermotoga maritima] |
| 2OSX_A | 2.34e-12 | 71 | 262 | 70 | 297 | ChainA, Endoglycoceramidase II [Rhodococcus sp.] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DJ77 | 3.04e-30 | 57 | 267 | 13 | 210 | Endoglucanase C OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celC PE=3 SV=1 |
| P23340 | 3.04e-30 | 57 | 267 | 13 | 210 | Endoglucanase C307 OS=Clostridium sp. (strain F1) OX=1508 GN=celC307 PE=1 SV=1 |
| P0C2S3 | 2.45e-28 | 57 | 267 | 13 | 210 | Endoglucanase C OS=Acetivibrio thermocellus OX=1515 GN=celC PE=1 SV=1 |
| W8QRE4 | 1.46e-18 | 29 | 227 | 3 | 229 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| P16169 | 3.56e-12 | 70 | 267 | 33 | 212 | Cellodextrinase A OS=Ruminococcus flavefaciens OX=1265 GN=celA PE=3 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as SP
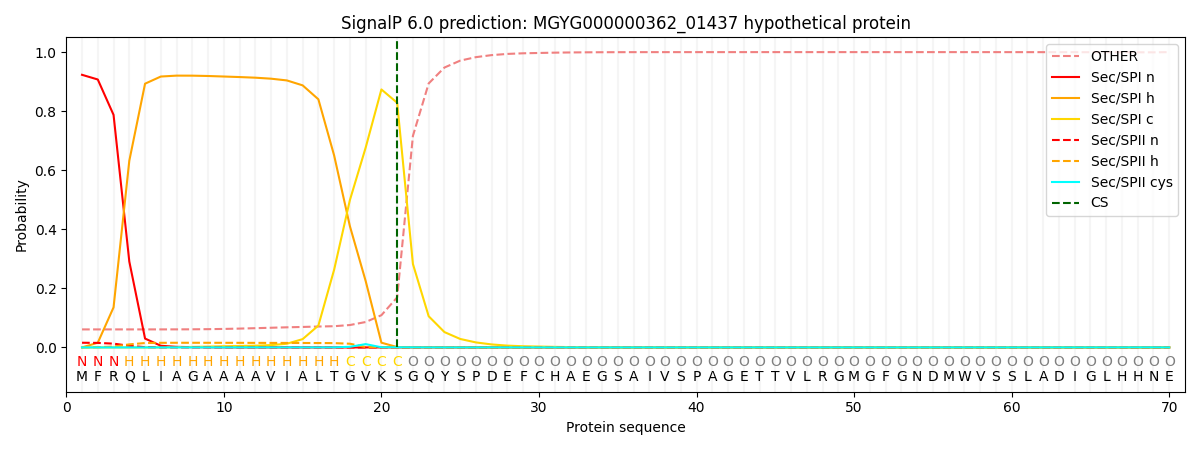
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.065398 | 0.916410 | 0.017178 | 0.000341 | 0.000317 | 0.000326 |
