You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000390_00138
You are here: Home > Sequence: MGYG000000390_00138
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; ; | |||||||||||
| CAZyme ID | MGYG000000390_00138 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12480; End: 15479 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 109 | 764 | 2.4e-65 | 0.6409574468085106 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 1.07e-23 | 91 | 762 | 20 | 679 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 4.75e-15 | 118 | 482 | 62 | 420 | beta-D-glucuronidase; Provisional |
| PRK10340 | ebgA | 1.45e-12 | 128 | 479 | 111 | 449 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam00703 | Glyco_hydro_2 | 1.83e-07 | 247 | 355 | 1 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| pfam02837 | Glyco_hydro_2_N | 4.64e-06 | 111 | 195 | 55 | 135 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALL53566.1 | 0.0 | 1 | 989 | 1 | 997 |
| QOV20252.1 | 0.0 | 4 | 997 | 3 | 1016 |
| AEI43263.1 | 7.84e-313 | 2 | 993 | 7 | 1010 |
| AFC30919.1 | 2.22e-312 | 2 | 993 | 7 | 1010 |
| AFH63244.2 | 4.46e-312 | 2 | 993 | 7 | 1010 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6ED2_A | 8.05e-17 | 129 | 495 | 94 | 454 | ChainA, Glycosyl hydrolase family 2, TIM barrel domain protein [Faecalibacterium duncaniae] |
| 7SF2_A | 2.20e-16 | 130 | 441 | 90 | 378 | ChainA, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_E Chain E, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_F Chain F, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838] |
| 6U7I_A | 1.19e-15 | 129 | 498 | 67 | 431 | Faecalibacteriumprausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_B Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_C Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii],6U7I_D Faecalibacterium prausnitzii Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 7KGY_A | 7.71e-14 | 129 | 502 | 72 | 439 | ChainA, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_B Chain B, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_C Chain C, Beta-glucuronidase [Faecalibacterium prausnitzii],7KGY_D Chain D, Beta-glucuronidase [Faecalibacterium prausnitzii] |
| 6LD0_A | 1.94e-13 | 126 | 480 | 67 | 482 | ChainA, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD0_B Chain B, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD0_C Chain C, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD0_D Chain D, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD6_A Chain A, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD6_B Chain B, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD6_C Chain C, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LD6_D Chain D, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDB_A Chain A, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDB_B Chain B, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDB_C Chain C, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDB_D Chain D, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDC_A Chain A, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDC_B Chain B, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDC_C Chain C, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDC_D Chain D, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDD_A Chain A, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDD_B Chain B, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDD_C Chain C, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1],6LDD_D Chain D, LacZ1 Beta-galactosidase [Bifidobacterium dentium Bd1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P77989 | 2.23e-13 | 107 | 481 | 28 | 390 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
| B4S2K9 | 4.70e-13 | 122 | 479 | 116 | 460 | Beta-galactosidase OS=Alteromonas mediterranea (strain DSM 17117 / CIP 110805 / LMG 28347 / Deep ecotype) OX=1774373 GN=lacZ PE=3 SV=1 |
| P06760 | 3.91e-09 | 140 | 589 | 114 | 542 | Beta-glucuronidase OS=Rattus norvegicus OX=10116 GN=Gusb PE=1 SV=1 |
| O18835 | 3.92e-09 | 140 | 589 | 114 | 545 | Beta-glucuronidase OS=Canis lupus familiaris OX=9615 GN=GUSB PE=1 SV=1 |
| P12265 | 5.15e-09 | 144 | 559 | 118 | 508 | Beta-glucuronidase OS=Mus musculus OX=10090 GN=Gusb PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
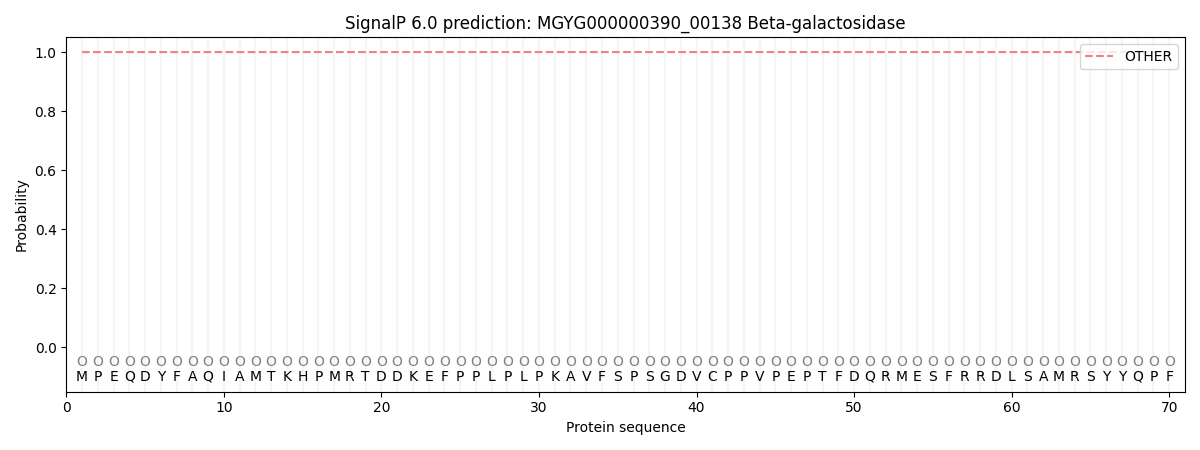
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000055 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
