You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000403_00712
You are here: Home > Sequence: MGYG000000403_00712
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA1381; UBA4716; | |||||||||||
| CAZyme ID | MGYG000000403_00712 | |||||||||||
| CAZy Family | GH167 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 127; End: 3237 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH167 | 299 | 847 | 4.5e-116 | 0.8047419804741981 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02449 | Glyco_hydro_42 | 3.59e-11 | 398 | 539 | 134 | 269 | Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. |
| COG1874 | GanA | 4.31e-06 | 415 | 495 | 173 | 256 | Beta-galactosidase GanA [Carbohydrate transport and metabolism]. |
| cd03143 | A4_beta-galactosidase_middle_domain | 0.002 | 653 | 763 | 12 | 120 | A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. A4 beta-galactosidase middle domain: a type 1 glutamine amidotransferase (GATase1)-like domain. This group includes proteins similar to beta-galactosidase from Thermus thermophilus. Beta-Galactosidase hydrolyzes the beta-1,4-D-galactosidic linkage of lactose, as well as those of related chromogens, o-nitrophenyl-beta-D-galactopyranoside (ONP-Gal) and 5-bromo-4-chloro-3-indolyl-beta-D-galactoside (X-gal). This A4 beta-galactosidase middle domain lacks the catalytic triad of typical GATase1 domains. The reactive Cys residue found in the sharp turn between a beta strand and an alpha helix termed the nucleophile elbow in typical GATase1 domains is not conserved in this group. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUL53851.1 | 1.07e-105 | 34 | 872 | 609 | 1494 |
| AIQ43288.1 | 8.09e-105 | 33 | 856 | 608 | 1475 |
| AIQ31958.1 | 1.43e-100 | 34 | 859 | 609 | 1479 |
| ALC81507.1 | 3.11e-99 | 21 | 830 | 347 | 1191 |
| BBI31589.1 | 2.87e-97 | 37 | 886 | 605 | 1481 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6PTM_A | 4.16e-31 | 324 | 767 | 213 | 682 | Crystalstructure of apo exo-carrageenase GH42 from Bacteroides ovatus [Bacteroides ovatus CL02T12C04] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
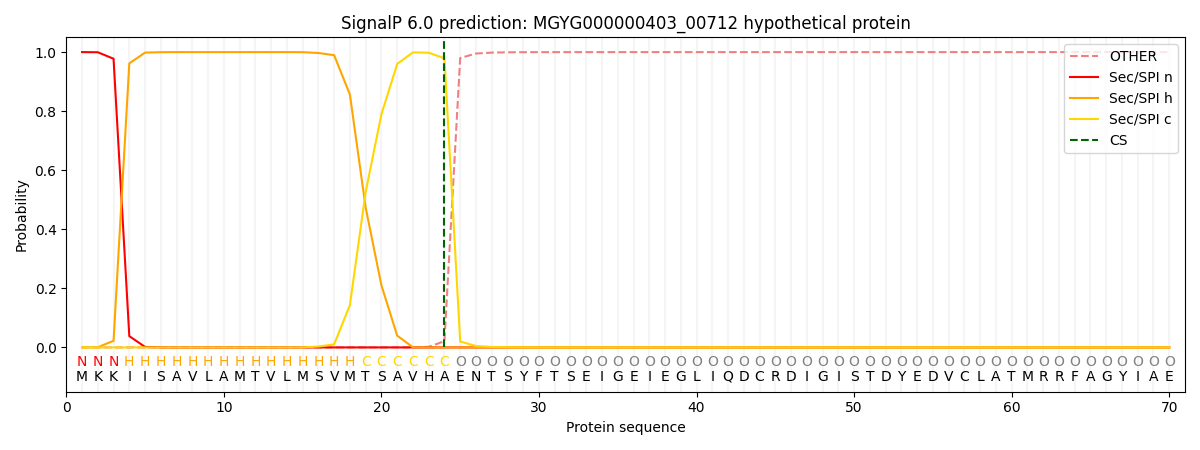
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000272 | 0.998976 | 0.000184 | 0.000179 | 0.000182 | 0.000168 |
