You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000414_01832
You are here: Home > Sequence: MGYG000000414_01832
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp900546065 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp900546065 | |||||||||||
| CAZyme ID | MGYG000000414_01832 | |||||||||||
| CAZy Family | CE15 | |||||||||||
| CAZyme Description | Carbohydrate esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 46049; End: 47272 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE15 | 25 | 407 | 2.3e-84 | 0.9925650557620818 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1506 | DAP2 | 2.63e-07 | 93 | 270 | 367 | 507 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QEC51453.1 | 5.17e-119 | 26 | 406 | 35 | 422 |
| QUT50761.1 | 4.62e-114 | 25 | 407 | 37 | 411 |
| QDV36691.1 | 9.04e-112 | 23 | 405 | 40 | 417 |
| SCD19288.1 | 6.60e-110 | 26 | 407 | 34 | 416 |
| QEG34875.1 | 8.51e-109 | 25 | 404 | 37 | 416 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6GRW_A | 2.21e-97 | 22 | 407 | 3 | 398 | GlucuronoylEsterase from Opitutus terrae (Au derivative) [Opitutus terrae PB90-1],6GS0_A Native Glucuronoyl Esterase from Opitutus terrae [Opitutus terrae PB90-1] |
| 6SYU_A | 3.83e-97 | 22 | 407 | 21 | 416 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with xylobiose [Opitutus terrae],6T0I_A The wild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with the aldotetrauronic acid XUX [Opitutus terrae] |
| 6SYR_A | 8.76e-97 | 25 | 407 | 35 | 427 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with D-glucuronate [Opitutus terrae PB90-1] |
| 6SYV_A | 1.08e-96 | 22 | 407 | 21 | 416 | Theglucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with D-glucuronate [Opitutus terrae PB90-1],6SZO_A The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with D-galacturonate [Opitutus terrae PB90-1],6T0E_A The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with benzyl D-glucuronoate and D-glucuronate [Opitutus terrae PB90-1],6T0E_B The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with benzyl D-glucuronoate and D-glucuronate [Opitutus terrae PB90-1] |
| 6GRY_A | 2.93e-96 | 25 | 402 | 13 | 392 | GlucuronoylEsterase from Solibacter usitatus. [Candidatus Solibacter usitatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A0K2VM55 | 3.65e-88 | 25 | 403 | 41 | 424 | Carbohydrate esterase MZ0003 OS=Unknown prokaryotic organism OX=2725 GN=MZ0003 PE=1 SV=1 |
| B2ABS0 | 2.99e-20 | 212 | 327 | 258 | 383 | 4-O-methyl-glucuronoyl methylesterase OS=Podospora anserina (strain S / ATCC MYA-4624 / DSM 980 / FGSC 10383) OX=515849 GN=ge1 PE=1 SV=1 |
| Q7S1X0 | 2.49e-19 | 212 | 356 | 176 | 329 | 4-O-methyl-glucuronoyl methylesterase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=Cip2 PE=1 SV=2 |
| P0CT87 | 4.15e-19 | 212 | 356 | 250 | 403 | 4-O-methyl-glucuronoyl methylesterase 1 OS=Phanerochaete chrysosporium (strain RP-78 / ATCC MYA-4764 / FGSC 9002) OX=273507 GN=e_gw1.18.61.1 PE=1 SV=1 |
| P0CU53 | 5.11e-19 | 119 | 356 | 84 | 338 | 4-O-methyl-glucuronoyl methylesterase 1 OS=Wolfiporia cocos (strain MD-104) OX=742152 GN=WOLCODRAFT_23632 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
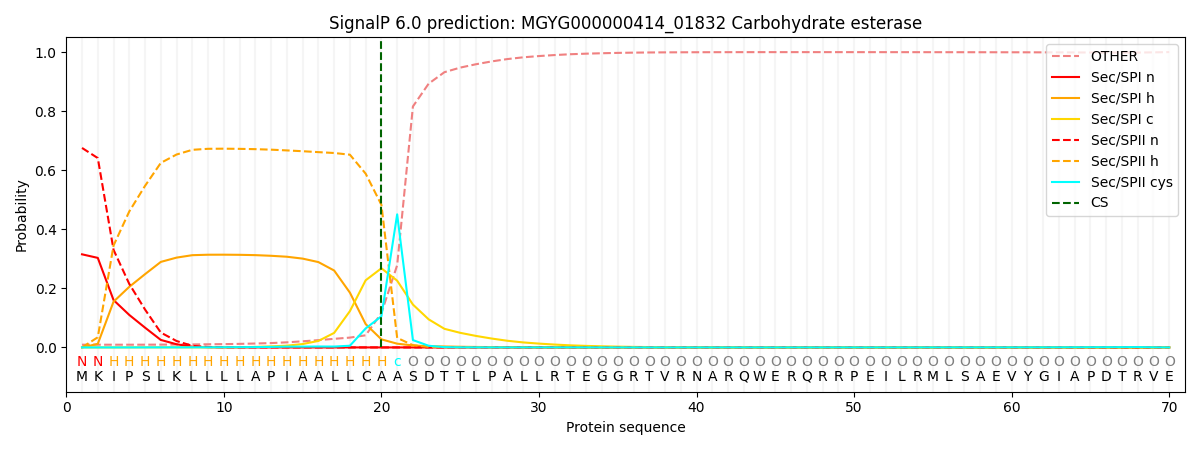
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.010818 | 0.307888 | 0.680713 | 0.000174 | 0.000178 | 0.000228 |
