You are browsing environment: HUMAN GUT
MGYG000000417_00438
Basic Information
help
Species
Acetatifactor sp900755865
Lineage
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Acetatifactor; Acetatifactor sp900755865
CAZyme ID
MGYG000000417_00438
CAZy Family
GH142
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000000417
2910146
MAG
Sweden
Europe
Gene Location
Start: 8419;
End: 10536
Strand: -
No EC number prediction in MGYG000000417_00438.
Family
Start
End
Evalue
family coverage
GH142
203
687
2.8e-56
0.9707724425887265
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam01204
Trehalase
1.02e-06
422
507
305
391
Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi.
more
PRK10137
PRK10137
4.34e-06
394
541
547
709
alpha-glucosidase; Provisional
more
COG3408
GDB1
4.56e-06
422
568
420
548
Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism].
more
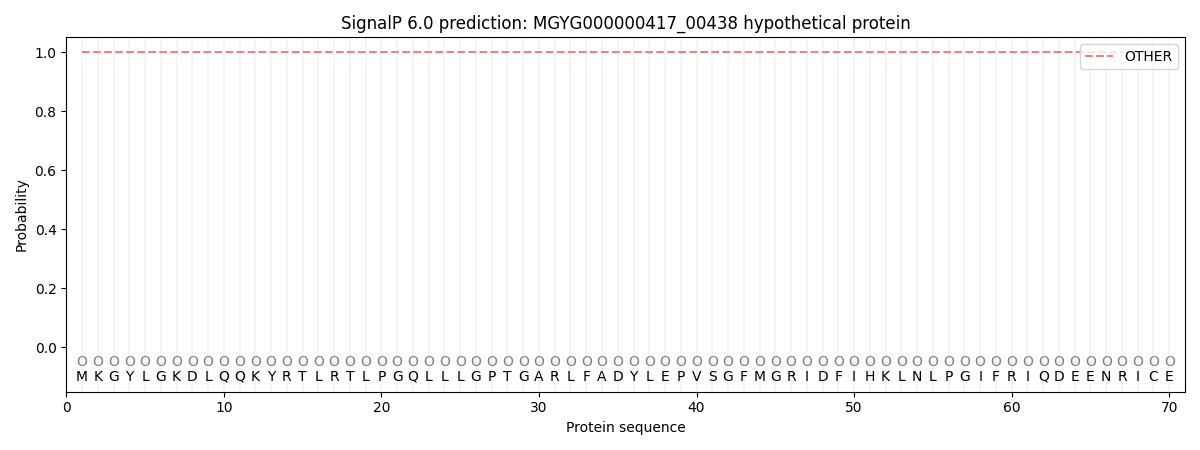
This protein is predicted as OTHER
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
1.000093
0.000000
0.000000
0.000000
0.000000
0.000000
There is no transmembrane helices in MGYG000000417_00438.