You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000426_01092
You are here: Home > Sequence: MGYG000000426_01092
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
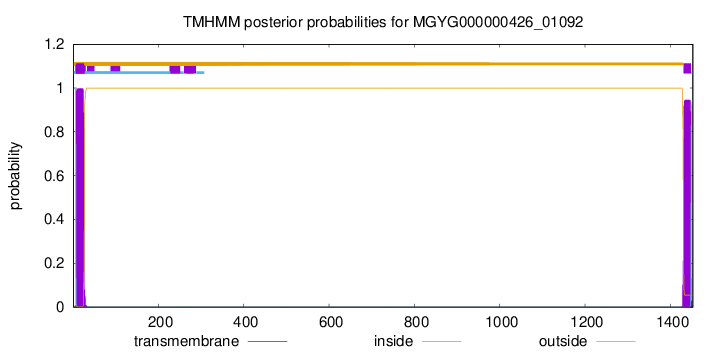
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; RF39; UBA660; UBA5026; | |||||||||||
| CAZyme ID | MGYG000000426_01092 | |||||||||||
| CAZy Family | CBM51 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7173; End: 11534 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM51 | 51 | 199 | 2.4e-28 | 0.9850746268656716 |
| CBM32 | 874 | 1002 | 1.1e-16 | 0.8870967741935484 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08305 | NPCBM | 4.07e-37 | 50 | 199 | 3 | 135 | NPCBM/NEW2 domain. This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. This domain has also been called the NEW2 domain (Naumoff DG. Phylogenetic analysis of alpha-galactosidases of the GH27 family. Molecular Biology (Engl Transl). (2004)38:388-399.) |
| smart00776 | NPCBM | 1.37e-23 | 50 | 199 | 5 | 144 | This novel putative carbohydrate binding module (NPCBM) domain is found at the N-terminus of glycosyl hydrolase family 98 proteins. |
| pfam00754 | F5_F8_type_C | 1.38e-09 | 886 | 1006 | 14 | 125 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam12396 | DUF3659 | 4.71e-04 | 1128 | 1161 | 8 | 52 | Protein of unknown function (DUF3659). This domain family is found in bacteria and eukaryotes, and is approximately 70 amino acids in length. |
| cd00057 | FA58C | 7.04e-04 | 874 | 969 | 19 | 109 | Substituted updates: Jan 31, 2002 |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCL58565.1 | 2.71e-260 | 36 | 1198 | 67 | 1191 |
| AMN35280.1 | 1.57e-163 | 3 | 1015 | 2 | 991 |
| AQW23377.1 | 4.25e-163 | 3 | 1015 | 2 | 991 |
| ATD49073.1 | 4.86e-163 | 3 | 1015 | 7 | 996 |
| VTS14484.1 | 5.42e-143 | 47 | 957 | 137 | 1046 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2VMH_A | 1.30e-14 | 47 | 199 | 6 | 150 | Thestructure of CBM51 from Clostridium perfringens GH95 [Clostridium perfringens] |
| 2VMG_A | 1.51e-14 | 47 | 199 | 12 | 156 | Thestructure of CBM51 from Clostridium perfringens GH95 in complex with methyl-galactose [Clostridium perfringens] |
| 2VMI_A | 3.29e-14 | 47 | 199 | 6 | 150 | Thestructure of seleno-methionine labelled CBM51 from Clostridium perfringens GH95 [Clostridium perfringens] |
| 7JRM_A | 1.92e-09 | 58 | 300 | 86 | 326 | ChainA, F5/8 type C domain protein [Clostridium perfringens ATCC 13124] |
| 7JRL_A | 2.03e-09 | 58 | 300 | 108 | 348 | ChainA, F5/8 type C domain protein [Clostridium perfringens ATCC 13124] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
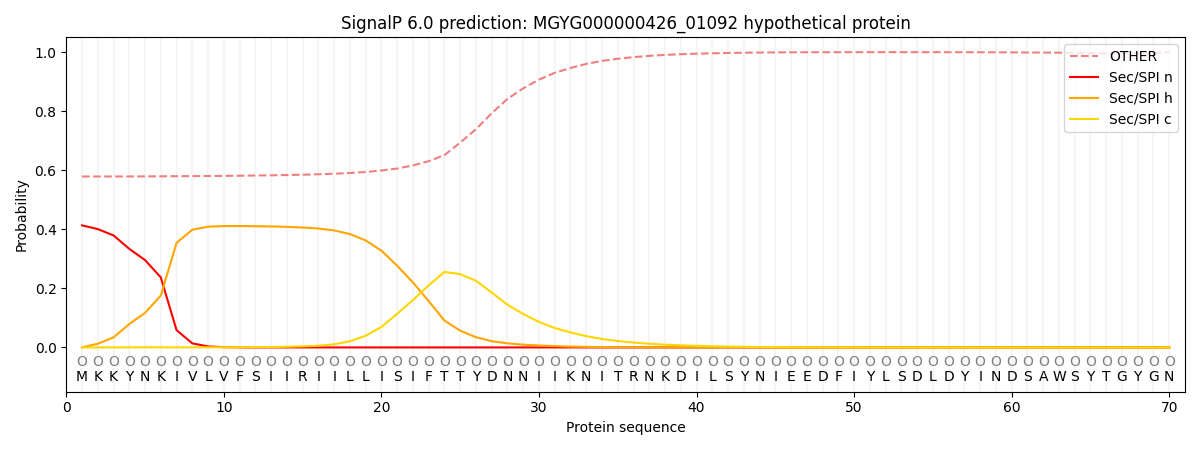
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.595186 | 0.392856 | 0.007475 | 0.000783 | 0.000471 | 0.003226 |