You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000431_00716
You are here: Home > Sequence: MGYG000000431_00716
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-288 sp000437395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; RFN20; CAG-288; CAG-288; CAG-288 sp000437395 | |||||||||||
| CAZyme ID | MGYG000000431_00716 | |||||||||||
| CAZy Family | GH84 | |||||||||||
| CAZyme Description | Hyaluronoglucosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 729212; End: 731902 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH84 | 175 | 477 | 3.1e-93 | 0.9864406779661017 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07555 | NAGidase | 6.93e-107 | 175 | 473 | 1 | 287 | beta-N-acetylglucosaminidase. This family has previously been described as a hyaluronidase. However, more recently it has been shown that this family has beta-N-acetylglucosaminidase activity. |
| pfam02838 | Glyco_hydro_20b | 1.71e-13 | 32 | 168 | 1 | 123 | Glycosyl hydrolase family 20, domain 2. This domain has a zincin-like fold. |
| pfam00754 | F5_F8_type_C | 1.49e-11 | 650 | 755 | 11 | 121 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam00754 | F5_F8_type_C | 6.15e-06 | 786 | 893 | 15 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SQG39213.1 | 9.32e-144 | 28 | 881 | 41 | 880 |
| ALG48841.1 | 1.60e-143 | 28 | 881 | 41 | 880 |
| QQA11067.1 | 2.24e-143 | 28 | 881 | 41 | 880 |
| ATD48581.1 | 2.33e-142 | 28 | 881 | 41 | 880 |
| AOY54048.1 | 2.33e-142 | 28 | 881 | 41 | 880 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6PWI_A | 2.05e-132 | 28 | 628 | 29 | 626 | Structureof CpGH84D [Clostridium perfringens ATCC 13124],6PWI_B Structure of CpGH84D [Clostridium perfringens ATCC 13124] |
| 6PV4_A | 1.43e-119 | 19 | 624 | 18 | 642 | Structureof CpGH84A [Clostridium perfringens ATCC 13124],6PV4_B Structure of CpGH84A [Clostridium perfringens ATCC 13124],6PV4_C Structure of CpGH84A [Clostridium perfringens ATCC 13124],6PV4_D Structure of CpGH84A [Clostridium perfringens ATCC 13124] |
| 6PV5_A | 2.32e-65 | 22 | 624 | 31 | 621 | Structureof CpGH84B [Clostridium perfringens ATCC 13124] |
| 2VVN_A | 5.30e-65 | 19 | 546 | 10 | 520 | BtGH84in complex with NH-Butylthiazoline [Bacteroides thetaiotaomicron VPI-5482],2VVN_B BtGH84 in complex with NH-Butylthiazoline [Bacteroides thetaiotaomicron VPI-5482],2VVS_A BtGH84 structure in complex with PUGNAc [Bacteroides thetaiotaomicron VPI-5482],2X0H_A BtGH84 Michaelis complex [Bacteroides thetaiotaomicron VPI-5482],2X0H_B BtGH84 Michaelis complex [Bacteroides thetaiotaomicron VPI-5482],4AIS_A A complex structure of BtGH84 [Bacteroides thetaiotaomicron VPI-5482],4AIS_B A complex structure of BtGH84 [Bacteroides thetaiotaomicron VPI-5482],7OU8_AAA Chain AAA, O-GlcNAcase BT_4395 [Bacteroides thetaiotaomicron VPI-5482],7OU8_BBB Chain BBB, O-GlcNAcase BT_4395 [Bacteroides thetaiotaomicron VPI-5482] |
| 5MI4_A | 1.15e-64 | 78 | 546 | 50 | 510 | BtGH84mutant with covalent modification by MA3 [Bacteroides thetaiotaomicron VPI-5482],5MI5_A BtGH84 mutant with covalent modification by MA3 in complex with PUGNAc [Bacteroides thetaiotaomicron VPI-5482],5MI6_A BtGH84 mutant with covalent modification by MA3 in complex with Thiamet G [Bacteroides thetaiotaomicron VPI-5482],5MI7_A BtGH84 mutant with covalent modification by MA4 in complex with PUGNAc [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P26831 | 2.08e-116 | 33 | 820 | 39 | 860 | Hyaluronoglucosaminidase OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagH PE=1 SV=2 |
| Q89ZI2 | 2.90e-64 | 19 | 546 | 10 | 520 | O-GlcNAcase BT_4395 OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_4395 PE=1 SV=1 |
| Q8XL08 | 2.39e-57 | 23 | 749 | 34 | 746 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagJ PE=1 SV=1 |
| Q0TR53 | 4.33e-57 | 23 | 749 | 34 | 746 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=nagJ PE=1 SV=1 |
| Q8VIJ5 | 5.91e-25 | 176 | 455 | 63 | 337 | Protein O-GlcNAcase OS=Rattus norvegicus OX=10116 GN=Oga PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
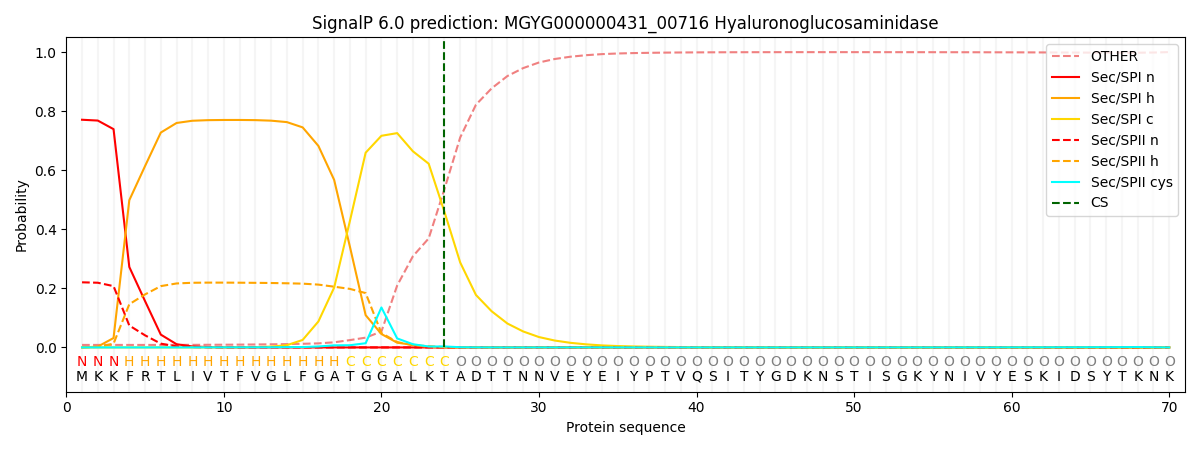
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.012766 | 0.759857 | 0.226094 | 0.000544 | 0.000360 | 0.000345 |
