You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000437_00867
You are here: Home > Sequence: MGYG000000437_00867
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp001941065 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp001941065 | |||||||||||
| CAZyme ID | MGYG000000437_00867 | |||||||||||
| CAZy Family | CE15 | |||||||||||
| CAZyme Description | Carbohydrate esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 33572; End: 34843 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE15 | 35 | 423 | 8.4e-105 | 0.9925650557620818 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03403 | PAF-AH_p_II | 0.009 | 251 | 285 | 218 | 252 | Platelet-activating factor acetylhydrolase, isoform II. Platelet-activating factor acetylhydrolase (PAF-AH) is a subfamily of phospholipases A2, responsible for inactivation of platelet-activating factor through cleavage of an acetyl group. Three known PAF-AHs are the brain heterotrimeric PAF-AH Ib, whose catalytic beta and gamma subunits are aligned in pfam02266, the extracellular, plasma PAF-AH (pPAF-AH), and the intracellular PAF-AH isoform II (PAF-AH II). This family aligns pPAF-AH and PAF-AH II, whose similarity was previously noted. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK63993.1 | 9.92e-273 | 5 | 423 | 6 | 426 |
| QDM10177.1 | 1.23e-195 | 23 | 423 | 290 | 685 |
| QUT79457.1 | 1.76e-195 | 23 | 423 | 301 | 696 |
| QUR45646.1 | 1.41e-194 | 23 | 423 | 290 | 685 |
| QDO70968.1 | 8.00e-190 | 16 | 420 | 31 | 431 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6SYR_A | 1.16e-144 | 19 | 423 | 19 | 427 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with D-glucuronate [Opitutus terrae PB90-1] |
| 6GRY_A | 5.61e-143 | 27 | 418 | 5 | 392 | GlucuronoylEsterase from Solibacter usitatus. [Candidatus Solibacter usitatus] |
| 6GRW_A | 1.38e-142 | 33 | 423 | 4 | 398 | GlucuronoylEsterase from Opitutus terrae (Au derivative) [Opitutus terrae PB90-1],6GS0_A Native Glucuronoyl Esterase from Opitutus terrae [Opitutus terrae PB90-1] |
| 6SYU_A | 2.54e-142 | 33 | 423 | 22 | 416 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with xylobiose [Opitutus terrae],6T0I_A The wild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with the aldotetrauronic acid XUX [Opitutus terrae] |
| 6SYV_A | 7.22e-142 | 33 | 423 | 22 | 416 | Theglucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with D-glucuronate [Opitutus terrae PB90-1],6SZO_A The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with D-galacturonate [Opitutus terrae PB90-1],6T0E_A The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with benzyl D-glucuronoate and D-glucuronate [Opitutus terrae PB90-1],6T0E_B The glucuronoyl esterase OtCE15A S267A variant from Opitutus terrae in complex with benzyl D-glucuronoate and D-glucuronate [Opitutus terrae PB90-1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A0K2VM55 | 2.31e-140 | 19 | 422 | 25 | 427 | Carbohydrate esterase MZ0003 OS=Unknown prokaryotic organism OX=2725 GN=MZ0003 PE=1 SV=1 |
| P0CU53 | 1.67e-22 | 29 | 380 | 35 | 344 | 4-O-methyl-glucuronoyl methylesterase 1 OS=Wolfiporia cocos (strain MD-104) OX=742152 GN=WOLCODRAFT_23632 PE=1 SV=1 |
| D8QLP9 | 4.72e-20 | 2 | 375 | 1 | 325 | 4-O-methyl-glucuronoyl methylesterase OS=Schizophyllum commune (strain H4-8 / FGSC 9210) OX=578458 GN=SCHCODRAFT_238770 PE=1 SV=1 |
| P0CT88 | 1.78e-19 | 15 | 375 | 28 | 337 | 4-O-methyl-glucuronoyl methylesterase 2 OS=Phanerochaete chrysosporium (strain RP-78 / ATCC MYA-4764 / FGSC 9002) OX=273507 GN=e_gw1.11.1537.1 PE=1 SV=1 |
| K5XDZ6 | 4.35e-19 | 36 | 375 | 44 | 337 | 4-O-methyl-glucuronoyl methylesterase OS=Phanerochaete carnosa (strain HHB-10118-sp) OX=650164 GN=PHACADRAFT_247750 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
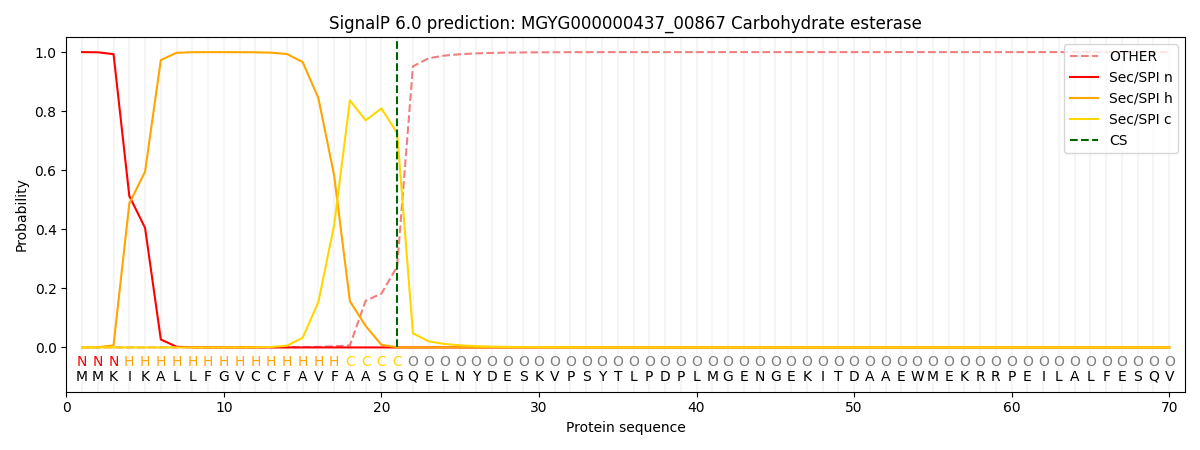
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000402 | 0.998786 | 0.000290 | 0.000171 | 0.000170 | 0.000157 |
