You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000441_00366
You are here: Home > Sequence: MGYG000000441_00366
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-312 sp900545705 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; CAG-312 sp900545705 | |||||||||||
| CAZyme ID | MGYG000000441_00366 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 171466; End: 173058 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 92 | 480 | 4.7e-40 | 0.9570957095709571 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 3.02e-30 | 168 | 480 | 15 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 3.47e-23 | 168 | 480 | 58 | 308 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 5.38e-21 | 168 | 486 | 81 | 343 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGA28189.1 | 2.02e-123 | 39 | 530 | 22 | 504 |
| AHF92621.1 | 5.06e-119 | 39 | 520 | 12 | 481 |
| AWI10666.1 | 8.74e-117 | 45 | 524 | 1 | 474 |
| AVM47074.1 | 1.08e-116 | 30 | 520 | 3 | 486 |
| QQZ02681.1 | 1.41e-112 | 38 | 520 | 18 | 485 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6FHF_A | 5.76e-14 | 145 | 480 | 41 | 358 | Highlyactive enzymes by automated modular backbone assembly and sequence design [Escherichia coli] |
| 6FHE_A | 2.82e-12 | 173 | 478 | 73 | 325 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
| 6LPS_A | 1.74e-11 | 171 | 482 | 68 | 351 | ChainA, Beta-xylanase [Halalkalibacterium halodurans] |
| 5EBA_A | 4.24e-11 | 171 | 482 | 68 | 355 | Crystalstructure of aromatic mutant (Y343A) of an alkali thermostable GH10 xylanase from Bacillus sp. NG-27 [Bacillus sp. NG-27] |
| 2UWF_A | 5.69e-11 | 171 | 482 | 67 | 350 | ChainA, ALKALINE ACTIVE ENDOXYLANASE [Halalkalibacterium halodurans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23556 | 1.61e-09 | 265 | 450 | 125 | 307 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynA PE=1 SV=1 |
| P07528 | 8.65e-09 | 171 | 482 | 113 | 396 | Endo-1,4-beta-xylanase A OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=xynA PE=1 SV=1 |
| P40942 | 3.33e-07 | 173 | 450 | 105 | 348 | Thermostable celloxylanase OS=Thermoclostridium stercorarium OX=1510 GN=xynB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
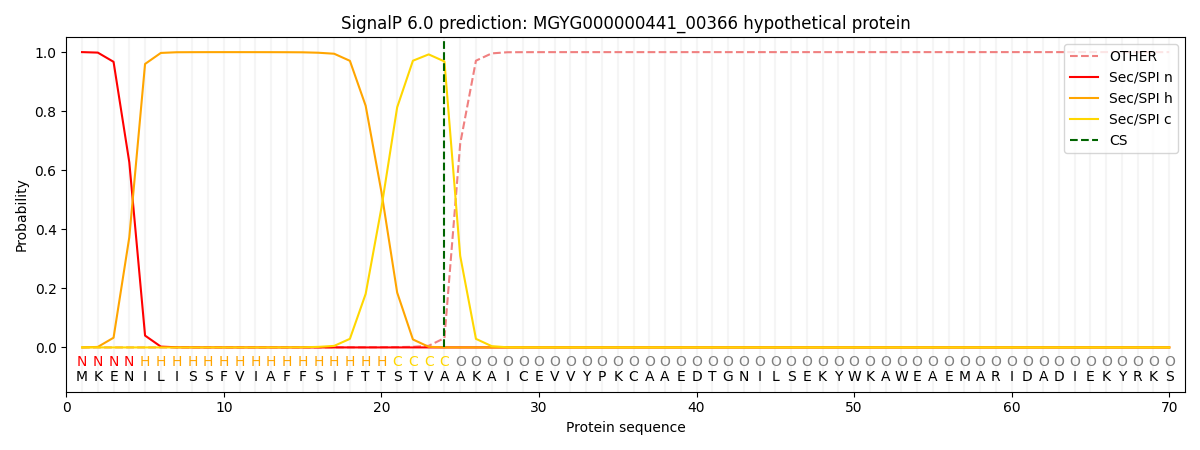
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000570 | 0.998692 | 0.000187 | 0.000175 | 0.000153 | 0.000150 |

