You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000470_00434
Basic Information
help
| Species |
Brachyspira pilosicoli
|
| Lineage |
Bacteria; Spirochaetota; Brachyspirae; Brachyspirales; Brachyspiraceae; Brachyspira; Brachyspira pilosicoli
|
| CAZyme ID |
MGYG000000470_00434
|
| CAZy Family |
GT0 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000470 |
2694868 |
MAG |
Fiji |
Oceania |
|
| Gene Location |
Start: 44872;
End: 45870
Strand: +
|
No EC number prediction in MGYG000000470_00434.
MGYG000000470_00434 has no CDD domain.
This protein is predicted as OTHER
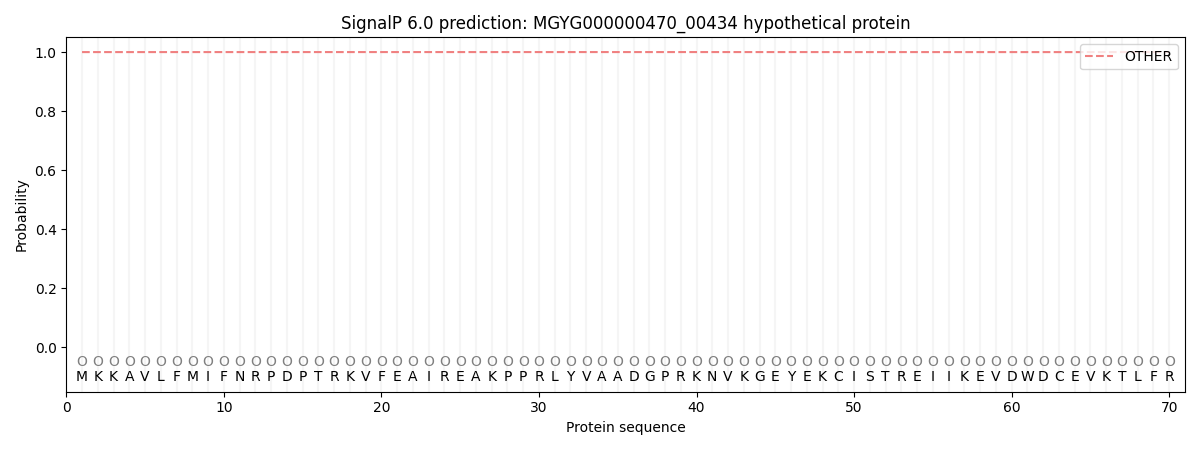
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000073
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000000470_00434.

