You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000480_01241
You are here: Home > Sequence: MGYG000000480_01241
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-312 sp900546565 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; CAG-312 sp900546565 | |||||||||||
| CAZyme ID | MGYG000000480_01241 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4769; End: 6376 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 82 | 486 | 1.8e-43 | 0.976897689768977 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 6.43e-30 | 150 | 485 | 3 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 2.63e-27 | 90 | 485 | 11 | 308 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 3.53e-24 | 162 | 491 | 79 | 343 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AHF92621.1 | 2.06e-163 | 38 | 527 | 17 | 483 |
| AVM47074.1 | 7.15e-163 | 23 | 535 | 2 | 496 |
| QGA28189.1 | 1.02e-160 | 29 | 535 | 18 | 504 |
| QQZ02681.1 | 3.30e-157 | 32 | 534 | 18 | 494 |
| AWI10666.1 | 1.72e-151 | 39 | 535 | 1 | 480 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6FHE_A | 4.18e-17 | 168 | 485 | 73 | 338 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
| 7D88_A | 6.36e-13 | 47 | 518 | 44 | 395 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
| 2DEP_A | 3.24e-12 | 168 | 485 | 66 | 340 | CrystalStructure of xylanase B from Clostridium stercorarium F9 [Thermoclostridium stercorarium],2DEP_B Crystal Structure of xylanase B from Clostridium stercorarium F9 [Thermoclostridium stercorarium] |
| 1VBR_A | 3.52e-12 | 152 | 492 | 43 | 323 | Crystalstructure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBR_B Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBU_A Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima],1VBU_B Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima] |
| 7CPK_A | 5.92e-12 | 162 | 365 | 65 | 229 | XylanaseR from Bacillus sp. TAR-1 [Bacillus sp. TAR1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P07528 | 3.94e-11 | 162 | 365 | 109 | 273 | Endo-1,4-beta-xylanase A OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=xynA PE=1 SV=1 |
| P40942 | 6.56e-10 | 168 | 485 | 105 | 379 | Thermostable celloxylanase OS=Thermoclostridium stercorarium OX=1510 GN=xynB PE=1 SV=1 |
| O69231 | 1.54e-09 | 166 | 485 | 63 | 328 | Endo-1,4-beta-xylanase B OS=Paenibacillus barcinonensis OX=198119 GN=xynB PE=1 SV=1 |
| P48789 | 7.89e-09 | 168 | 490 | 87 | 369 | Endo-1,4-beta-xylanase A OS=Prevotella ruminicola OX=839 GN=xynA PE=3 SV=1 |
| Q60041 | 2.96e-08 | 152 | 494 | 62 | 344 | Endo-1,4-beta-xylanase B OS=Thermotoga neapolitana OX=2337 GN=xynB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
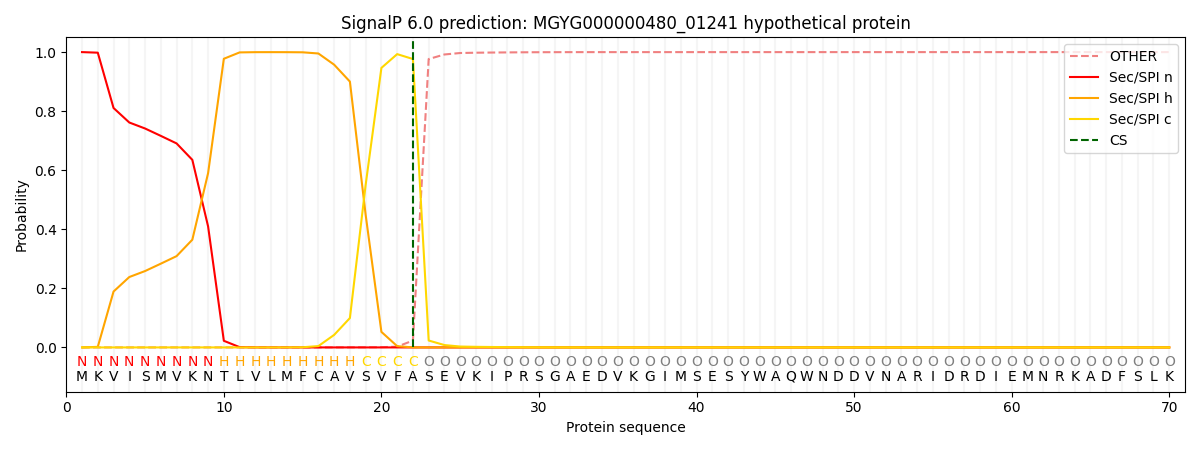
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000325 | 0.998975 | 0.000187 | 0.000169 | 0.000158 | 0.000147 |
