You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000480_01382
You are here: Home > Sequence: MGYG000000480_01382
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-312 sp900546565 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; CAG-312 sp900546565 | |||||||||||
| CAZyme ID | MGYG000000480_01382 | |||||||||||
| CAZy Family | GH109 | |||||||||||
| CAZyme Description | Inositol 2-dehydrogenase/D-chiro-inositol 3-dehydrogenase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8920; End: 10260 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 31 | 206 | 1.1e-19 | 0.39849624060150374 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0673 | MviM | 7.81e-35 | 32 | 408 | 3 | 337 | Predicted dehydrogenase [General function prediction only]. |
| pfam01408 | GFO_IDH_MocA | 1.06e-14 | 33 | 169 | 1 | 115 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| TIGR04380 | myo_inos_iolG | 2.72e-13 | 32 | 200 | 1 | 145 | inositol 2-dehydrogenase. All members of the seed alignment for this model are known or predicted inositol 2-dehydrogenase sequences co-clustered with other enzymes for catabolism of myo-inositol or closely related compounds. Inositol 2-dehydrogenase catalyzes the first step in inositol catabolism. Members of this family may vary somewhat in their ranges of acceptable substrates and some may act on analogs to myo-inositol rather than myo-inositol per se. [Energy metabolism, Sugars] |
| PRK11579 | PRK11579 | 3.11e-08 | 95 | 201 | 39 | 146 | putative oxidoreductase; Provisional |
| PRK10206 | PRK10206 | 6.94e-08 | 108 | 240 | 56 | 197 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AQT66975.1 | 2.92e-104 | 2 | 446 | 12 | 425 |
| AOS46431.1 | 1.46e-100 | 2 | 446 | 10 | 432 |
| AWI08226.1 | 3.46e-98 | 1 | 446 | 4 | 430 |
| QXD26316.1 | 4.36e-96 | 2 | 446 | 9 | 430 |
| QXD30383.1 | 4.36e-96 | 2 | 446 | 9 | 430 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3HNP_A | 4.10e-11 | 108 | 241 | 57 | 199 | CrystalStructure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579],3HNP_B Crystal Structure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579],3HNP_C Crystal Structure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579],3HNP_D Crystal Structure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579],3HNP_E Crystal Structure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579],3HNP_F Crystal Structure of an Oxidoreductase from Bacillus cereus. Northeast Structural Genomics Consortium target id BcR251 [Bacillus cereus ATCC 14579] |
| 6O15_A | 1.04e-08 | 87 | 282 | 34 | 197 | Crystalstructure of a putative oxidoreductase YjhC from Escherichia coli in complex with NAD(H) [Escherichia coli K-12],6O15_B Crystal structure of a putative oxidoreductase YjhC from Escherichia coli in complex with NAD(H) [Escherichia coli K-12] |
| 3GDO_A | 1.72e-07 | 108 | 259 | 57 | 188 | Crystalstructure of putative oxidoreductase yvaA from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168],3GDO_B Crystal structure of putative oxidoreductase yvaA from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168] |
| 3GFG_A | 1.77e-07 | 108 | 259 | 66 | 197 | Structureof putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_B Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_C Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_D Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_E Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_F Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_G Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_H Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_I Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_J Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_K Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168],3GFG_L Structure of putative oxidoreductase yvaA from Bacillus subtilis in triclinic form [Bacillus subtilis subsp. subtilis str. 168] |
| 3F4L_A | 2.18e-07 | 108 | 241 | 57 | 199 | Crystalstructure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12],3F4L_B Crystal structure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12],3F4L_C Crystal structure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12],3F4L_D Crystal structure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12],3F4L_E Crystal structure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12],3F4L_F Crystal structure of a probable oxidoreductase yhhX in Triclinic form. Northeast Structural Genomics target ER647 [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q01S58 | 3.90e-08 | 2 | 208 | 5 | 204 | Glycosyl hydrolase family 109 protein OS=Solibacter usitatus (strain Ellin6076) OX=234267 GN=Acid_6590 PE=3 SV=1 |
| A8H2K3 | 5.37e-08 | 31 | 204 | 51 | 208 | Glycosyl hydrolase family 109 protein OS=Shewanella pealeana (strain ATCC 700345 / ANG-SQ1) OX=398579 GN=Spea_1465 PE=3 SV=1 |
| P39353 | 5.71e-08 | 87 | 282 | 34 | 197 | Uncharacterized oxidoreductase YjhC OS=Escherichia coli (strain K12) OX=83333 GN=yjhC PE=1 SV=2 |
| O42896 | 9.97e-08 | 105 | 258 | 57 | 191 | Uncharacterized oxidoreductase C115.03 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC115.03 PE=3 SV=1 |
| A1S4U5 | 3.86e-07 | 1 | 204 | 4 | 208 | Glycosyl hydrolase family 109 protein OS=Shewanella amazonensis (strain ATCC BAA-1098 / SB2B) OX=326297 GN=Sama_1194 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as TAT
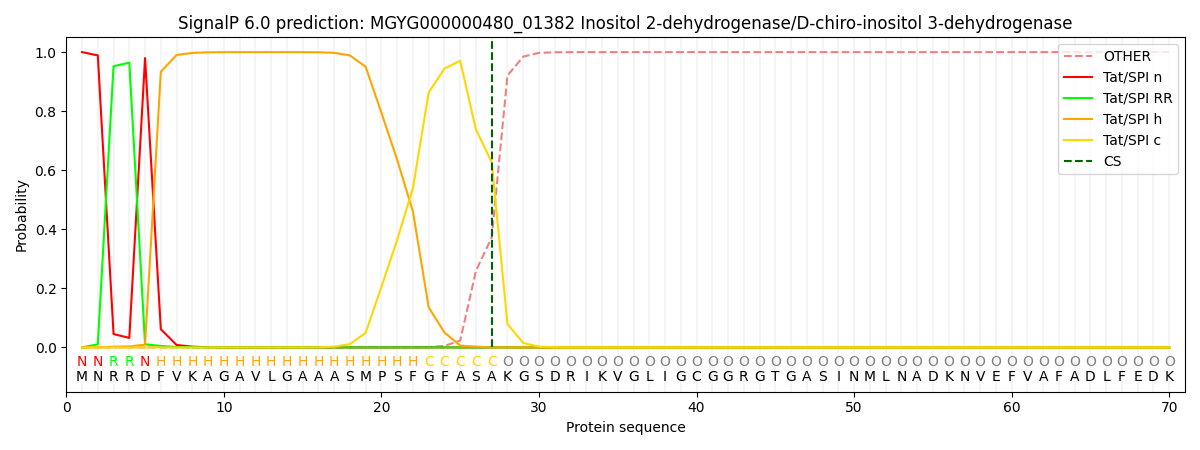
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 0.000000 | 0.999971 | 0.000000 | 0.000000 |
