You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000504_00831
You are here: Home > Sequence: MGYG000000504_00831
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
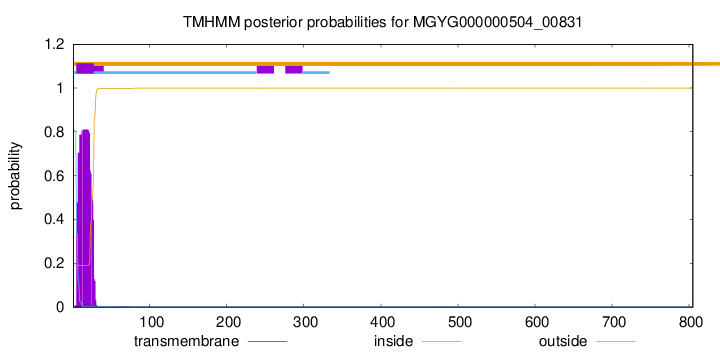
TMHMM annotations
Basic Information help
| Species | CAG-312 sp000438015 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312; CAG-312 sp000438015 | |||||||||||
| CAZyme ID | MGYG000000504_00831 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 47213; End: 49630 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 548 | 804 | 1e-57 | 0.9956521739130435 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam12831 | FAD_oxidored | 2.78e-115 | 57 | 531 | 1 | 420 | FAD dependent oxidoreductase. This family of proteins contains FAD dependent oxidoreductases and related proteins. |
| cd08023 | GH16_laminarinase_like | 1.47e-72 | 548 | 804 | 1 | 235 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| cd08024 | GH16_CCF | 1.31e-31 | 546 | 778 | 1 | 273 | Coelomic cytolytic factor, member of glycosyl hydrolase family 16. Subgroup of glucanases of unknown function that are related to beta-GRP (beta-1,3-glucan recognition protein), but contain active site residues. Beta-GRPs are one group of pattern recognition receptors (PRRs), also referred to as biosensor proteins, that complexes with pathogen-associated beta-1,3-glucans and then transduces signals necessary for activation of an appropriate innate immune response. Beta-GRPs are present in insects and lack all catalytic residues. This subgroup contains related proteins that still contain the active site and are widely distributed in eukaryotes. Their structures adopt a jelly roll fold with a deep active site channel harboring the catalytic residues, like those of other glycosyl hydrolase family 16 members. |
| cd02182 | GH16_Strep_laminarinase_like | 2.60e-23 | 545 | 805 | 3 | 259 | Streptomyces laminarinase-like, member of glycosyl hydrolase family 16. Proteins similar to Streptomyces sioyaensis beta-1,3-glucanase (laminarinase) present in Actinomycetales as well as Peziomycotina. Laminarinases belong to glycosyl hydrolase family 16 and hydrolyze the glycosidic bond of the 1,3-beta-linked glucan, a major component of fungal and plant cell walls and the structural and storage polysaccharides (laminarin) of marine macro-algae. Members of the GH16 family have a conserved jelly roll fold with an active site channel. |
| cd00413 | Glyco_hydrolase_16 | 6.00e-22 | 550 | 804 | 1 | 210 | glycosyl hydrolase family 16. The O-Glycosyl hydrolases are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A glycosyl hydrolase classification system based on sequence similarity has led to the definition of more than 95 different families inlcuding glycosyl hydrolase family 16. Family 16 includes lichenase, xyloglucan endotransglycosylase (XET), beta-agarase, kappa-carrageenase, endo-beta-1,3-glucanase, endo-beta-1,3-1,4-glucanase, and endo-beta-galactosidase, all of which have a conserved jelly roll fold with a deep active site channel harboring the catalytic residues. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ANU62583.2 | 7.79e-112 | 545 | 805 | 356 | 610 |
| ASB36926.1 | 7.79e-112 | 545 | 805 | 356 | 610 |
| QQR10084.1 | 1.61e-111 | 545 | 805 | 382 | 636 |
| QDO67480.1 | 7.73e-111 | 545 | 805 | 26 | 281 |
| QUU07365.1 | 3.51e-109 | 545 | 805 | 29 | 284 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6T2N_AAA | 2.81e-49 | 535 | 805 | 39 | 303 | ChainAAA, Glycoside hydrolase family 16 protein [Akkermansia muciniphila],6T2N_BBB Chain BBB, Glycoside hydrolase family 16 protein [Akkermansia muciniphila] |
| 4DFS_A | 9.64e-42 | 545 | 805 | 19 | 263 | Structureof the catalytic domain of an endo-1,3-beta-glucanase (laminarinase) from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],4DFS_B Structure of the catalytic domain of an endo-1,3-beta-glucanase (laminarinase) from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1] |
| 3AZX_A | 1.51e-41 | 545 | 805 | 11 | 255 | Crystalstructure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZX_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZY_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3AZZ_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3AZZ_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with gluconolactone [Thermotoga maritima MSB8],3B00_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B00_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 in complex with cetyltrimethylammonium bromide [Thermotoga maritima MSB8],3B01_A Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_B Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_C Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8],3B01_D Crystal structure of the laminarinase catalytic domain from Thermotoga maritima MSB8 [Thermotoga maritima MSB8] |
| 4BQ1_A | 9.65e-37 | 546 | 805 | 13 | 256 | Crystalstructure of of LamAcat from Zobellia galactanivorans [Zobellia galactanivorans],4BQ1_B Crystal structure of of LamAcat from Zobellia galactanivorans [Zobellia galactanivorans] |
| 6XOF_A | 1.35e-36 | 541 | 805 | 21 | 267 | ChainA, GH16 family protein [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P45798 | 1.33e-34 | 545 | 805 | 41 | 283 | Beta-glucanase OS=Rhodothermus marinus OX=29549 GN=bglA PE=1 SV=1 |
| P23903 | 3.99e-26 | 544 | 805 | 423 | 680 | Glucan endo-1,3-beta-glucosidase A1 OS=Niallia circulans OX=1397 GN=glcA PE=1 SV=1 |
| Q27082 | 5.72e-23 | 543 | 805 | 24 | 253 | Clotting factor G alpha subunit OS=Tachypleus tridentatus OX=6853 PE=1 SV=1 |
| C1IE32 | 6.33e-20 | 546 | 796 | 22 | 263 | Glucan endo-1,3-beta-glucosidase OS=Cryptopygus antarcticus OX=187623 PE=1 SV=1 |
| Q9ZG90 | 9.47e-19 | 518 | 805 | 32 | 288 | Keratan-sulfate endo-1,4-beta-galactosidase OS=Sphingobacterium multivorum OX=28454 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000254 | 0.999083 | 0.000164 | 0.000176 | 0.000163 | 0.000146 |