You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000514_00602
You are here: Home > Sequence: MGYG000000514_00602
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
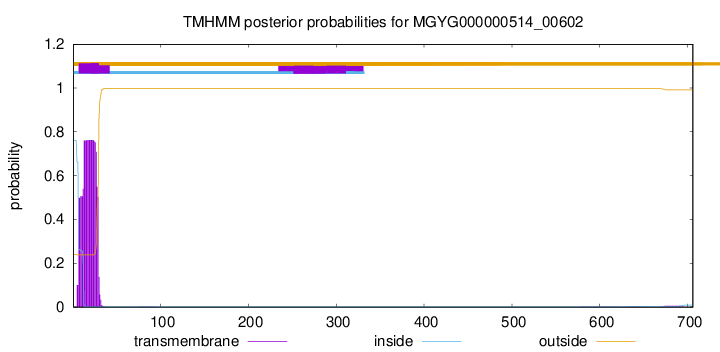
TMHMM annotations
Basic Information help
| Species | CAG-590 sp900769115 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; CAG-590; CAG-590 sp900769115 | |||||||||||
| CAZyme ID | MGYG000000514_00602 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10015; End: 12135 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4193 | LytD | 7.30e-19 | 366 | 597 | 43 | 245 | Beta- N-acetylglucosaminidase [Carbohydrate transport and metabolism]. |
| COG3103 | YgiM | 4.60e-11 | 130 | 265 | 36 | 153 | Uncharacterized conserved protein YgiM, contains N-terminal SH3 domain, DUF1202 family [General function prediction only]. |
| pfam08239 | SH3_3 | 3.47e-09 | 207 | 262 | 1 | 54 | Bacterial SH3 domain. |
| COG3103 | YgiM | 9.96e-06 | 55 | 185 | 35 | 150 | Uncharacterized conserved protein YgiM, contains N-terminal SH3 domain, DUF1202 family [General function prediction only]. |
| pfam08239 | SH3_3 | 1.69e-05 | 128 | 185 | 3 | 54 | Bacterial SH3 domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK83300.1 | 6.93e-110 | 82 | 705 | 4 | 631 |
| QWT53775.1 | 3.04e-107 | 239 | 705 | 79 | 528 |
| QNM00733.1 | 3.53e-105 | 239 | 705 | 79 | 528 |
| AEN96074.1 | 1.76e-104 | 133 | 705 | 51 | 611 |
| CUH91961.1 | 3.44e-103 | 54 | 706 | 149 | 912 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4PI7_A | 8.51e-10 | 443 | 584 | 87 | 217 | ChainA, Autolysin E [Staphylococcus aureus subsp. aureus Mu50],4PI9_A Chain A, Autolysin E [Staphylococcus aureus subsp. aureus Mu50],4PIA_A Chain A, Autolysin E [Staphylococcus aureus subsp. aureus Mu50] |
| 2KYB_A | 3.16e-09 | 197 | 261 | 2 | 60 | Solutionstructure of CpR82G from Clostridium perfringens. North East Structural Genomics Consortium Target CpR82g [Clostridium perfringens ATCC 13124] |
| 4PI8_A | 5.11e-09 | 443 | 584 | 87 | 217 | ChainA, Autolysin E [Staphylococcus aureus subsp. aureus Mu50] |
| 2KT8_A | 6.57e-09 | 197 | 261 | 3 | 61 | SolutionNMR structure of the CPE1231(468-535) protein from Clostridium perfringens, Northeast Structural Genomics Consortium Target CpR82B [Clostridium perfringens] |
| 6FXO_A | 6.67e-08 | 371 | 596 | 37 | 243 | ChainA, Bifunctional autolysin [Staphylococcus aureus subsp. aureus Mu50] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8CPQ1 | 1.84e-09 | 371 | 596 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200) OX=176280 GN=atl PE=3 SV=1 |
| O33635 | 4.16e-09 | 371 | 596 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis OX=1282 GN=atl PE=1 SV=1 |
| Q5HQB9 | 4.16e-09 | 371 | 596 | 1128 | 1334 | Bifunctional autolysin OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=atl PE=3 SV=1 |
| O32041 | 1.55e-06 | 50 | 279 | 35 | 257 | Putative N-acetylmuramoyl-L-alanine amidase YrvJ OS=Bacillus subtilis (strain 168) OX=224308 GN=yrvJ PE=3 SV=1 |
| Q931U5 | 1.66e-06 | 371 | 596 | 1041 | 1247 | Bifunctional autolysin OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=atl PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.003010 | 0.995786 | 0.000523 | 0.000234 | 0.000198 | 0.000208 |