You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000526_00079
You are here: Home > Sequence: MGYG000000526_00079
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900770395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900770395 | |||||||||||
| CAZyme ID | MGYG000000526_00079 | |||||||||||
| CAZy Family | CE19 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 105351; End: 106502 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE19 | 58 | 383 | 1.9e-27 | 0.8463855421686747 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0412 | DLH | 2.21e-12 | 111 | 383 | 13 | 234 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam12715 | Abhydrolase_7 | 9.32e-11 | 56 | 341 | 45 | 326 | Abhydrolase family. This is a family of probable bacterial abhydrolases. |
| COG1506 | DAP2 | 1.91e-10 | 92 | 383 | 358 | 617 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam00326 | Peptidase_S9 | 3.33e-09 | 175 | 383 | 9 | 210 | Prolyl oligopeptidase family. |
| COG3458 | Axe1 | 4.18e-06 | 176 | 340 | 105 | 282 | Cephalosporin-C deacetylase or related acetyl esterase [Secondary metabolites biosynthesis, transport and catabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY37157.1 | 4.87e-125 | 15 | 381 | 361 | 726 |
| QEG26086.1 | 1.03e-19 | 85 | 383 | 50 | 329 |
| VTS00541.1 | 1.03e-19 | 85 | 383 | 50 | 329 |
| AWM40652.1 | 1.03e-19 | 85 | 383 | 50 | 329 |
| AMV40306.1 | 2.14e-19 | 76 | 383 | 76 | 362 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34973 | 1.02e-26 | 79 | 354 | 17 | 270 | Putative hydrolase YtaP OS=Bacillus subtilis (strain 168) OX=224308 GN=ytaP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
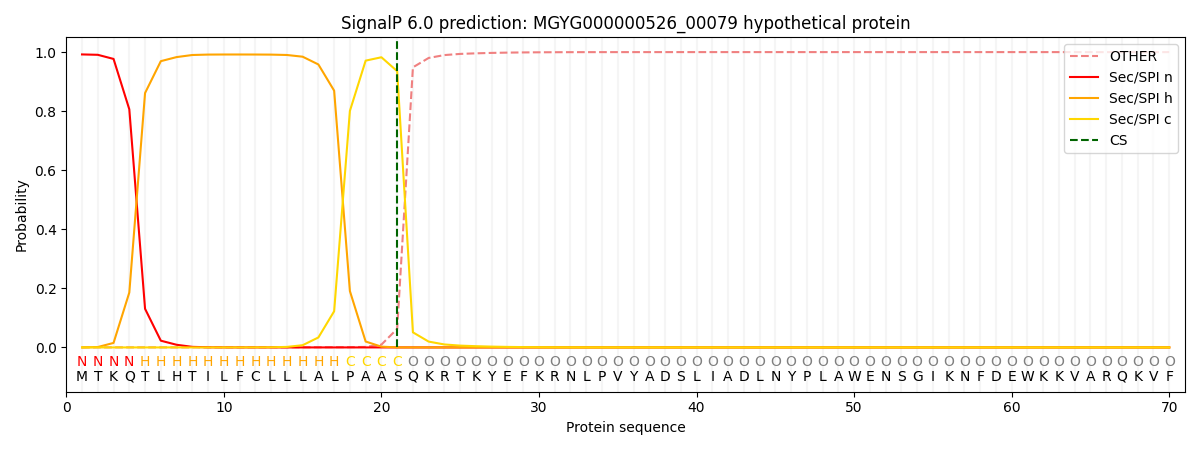
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001065 | 0.989826 | 0.008445 | 0.000249 | 0.000214 | 0.000200 |
