You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000526_00583
You are here: Home > Sequence: MGYG000000526_00583
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900770395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900770395 | |||||||||||
| CAZyme ID | MGYG000000526_00583 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 100922; End: 103369 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE6 | 582 | 692 | 3.5e-31 | 0.9797979797979798 |
| CE1 | 34 | 227 | 1.3e-24 | 0.6960352422907489 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3509 | LpqC | 2.31e-39 | 1 | 272 | 8 | 307 | Poly(3-hydroxybutyrate) depolymerase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam03629 | SASA | 1.40e-27 | 518 | 766 | 1 | 226 | Carbohydrate esterase, sialic acid-specific acetylesterase. The catalytic triad of this esterase enzyme comprises residues Ser127, His403 and Asp391 in UniProtKB:P70665. |
| TIGR01840 | esterase_phb | 2.56e-20 | 39 | 201 | 1 | 194 | esterase, PHB depolymerase family. This model describes a subfamily among lipases of the ab-hydrolase family. This subfamily includes bacterial depolymerases for poly(3-hydroxybutyrate) (PHB) and related polyhydroxyalkanoates (PHA), as well as acetyl xylan esterases, feruloyl esterases, and others from fungi. [Fatty acid and phospholipid metabolism, Degradation] |
| COG4099 | COG4099 | 3.23e-11 | 40 | 193 | 177 | 333 | Predicted peptidase [General function prediction only]. |
| pfam02230 | Abhydrolase_2 | 4.96e-09 | 41 | 192 | 6 | 173 | Phospholipase/Carboxylesterase. This family consists of both phospholipases and carboxylesterases with broad substrate specificity, and is structurally related to alpha/beta hydrolases pfam00561. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIA08072.1 | 1.02e-96 | 498 | 768 | 3 | 276 |
| QNT67597.1 | 2.54e-94 | 504 | 780 | 18 | 296 |
| QNL40592.1 | 1.02e-93 | 516 | 789 | 21 | 294 |
| ALJ48336.1 | 1.05e-92 | 516 | 785 | 21 | 289 |
| QRQ55180.1 | 1.05e-92 | 516 | 785 | 21 | 289 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4MGQ_A | 3.05e-15 | 274 | 427 | 5 | 146 | PbXyn10CCBM APO [Prevotella bryantii B14] |
| 3WYD_A | 2.71e-08 | 52 | 204 | 38 | 183 | C-terminalesterase domain of LC-Est1 [uncultured organism],3WYD_B C-terminal esterase domain of LC-Est1 [uncultured organism] |
| 3DOH_A | 6.28e-08 | 41 | 202 | 162 | 335 | CrystalStructure of a Thermostable Esterase [Thermotoga maritima],3DOH_B Crystal Structure of a Thermostable Esterase [Thermotoga maritima],3DOI_A Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima],3DOI_B Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima] |
| 5X6S_A | 8.23e-07 | 35 | 166 | 15 | 148 | Acetylxylan esterase from Aspergillus awamori [Aspergillus awamori],5X6S_B Acetyl xylan esterase from Aspergillus awamori [Aspergillus awamori] |
| 1ZMB_A | 2.89e-06 | 522 | 762 | 5 | 222 | CrystalStructure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_B Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_C Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_D Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_E Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824],1ZMB_F Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 [Clostridium acetobutylicum ATCC 824] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EXZ4 | 2.05e-80 | 495 | 762 | 24 | 289 | Carbohydrate acetyl esterase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe1-6A PE=1 SV=1 |
| B8YG19 | 4.64e-51 | 34 | 274 | 55 | 282 | Bifunctional acetylxylan esterase/xylanase XynS20E OS=Neocallimastix patriciarum OX=4758 GN=xynS20E PE=1 SV=1 |
| Q9Y871 | 1.23e-14 | 37 | 274 | 292 | 535 | Feruloyl esterase B OS=Piromyces equi OX=99929 GN=ESTA PE=2 SV=1 |
| P52090 | 2.78e-12 | 1 | 162 | 16 | 179 | Poly(3-hydroxyalkanoate) depolymerase C OS=Paucimonas lemoignei OX=29443 GN=phaZ1 PE=3 SV=1 |
| G2QND5 | 7.99e-11 | 25 | 175 | 19 | 175 | Feruloyl esterase B OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=Fae1a PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
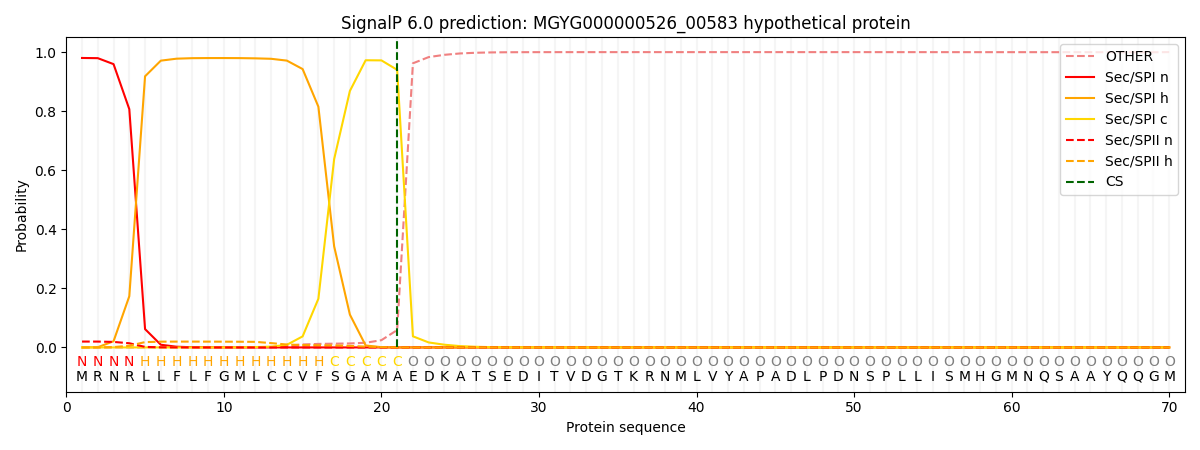
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000537 | 0.975375 | 0.023293 | 0.000290 | 0.000240 | 0.000215 |
