You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000529_00921
You are here: Home > Sequence: MGYG000000529_00921
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1585 sp900553205 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Cyanobacteria; Vampirovibrionia; Gastranaerophilales; Gastranaerophilaceae; UMGS1585; UMGS1585 sp900553205 | |||||||||||
| CAZyme ID | MGYG000000529_00921 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 114251; End: 115912 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 16 | 492 | 2.9e-101 | 0.8740740740740741 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1807 | ArnT | 2.24e-46 | 1 | 445 | 1 | 431 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| PRK13279 | arnT | 3.85e-37 | 6 | 430 | 3 | 423 | lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
| pfam13231 | PMT_2 | 1.25e-16 | 67 | 230 | 1 | 157 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
| pfam02366 | PMT | 4.46e-12 | 21 | 232 | 12 | 232 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This is a family of Dolichyl-phosphate-mannose-protein mannosyltransferase proteins EC:2.4.1.109. These proteins are responsible for O-linked glycosylation of proteins, they catalyze the reaction:- Dolichyl phosphate D-mannose + protein <=> dolichyl phosphate + O-D-mannosyl-protein. Also in this family is Drosophila rotated abdomen protein which is a putative mannosyltransferase. This family appears to be distantly related to pfam02516 (A Bateman pers. obs.). This family also contains sequences from ArnTs (4-amino-4-deoxy-L-arabinose lipid A transferase). They catalyze the addition of 4-amino-4-deoxy-l-arabinose (l-Ara4N) to the lipid A moiety of the lipopolysaccharide. This is a critical modification enabling bacteria (e.g. Escherichia coli and Salmonella typhimurium) to resist killing by antimicrobial peptides such as polymyxins. Members such as undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase are predicted to have 12 trans-membrane regions. The N-terminal portion of these proteins is hypothesized to have a conserved glycosylation activity which is shared between distantly related oligosaccharyltransferases ArnT and PglB families. |
| COG4346 | COG4346 | 7.44e-04 | 92 | 361 | 158 | 425 | Predicted membrane-bound dolichyl-phosphate-mannose-protein mannosyltransferase [Posttranslational modification, protein turnover, chaperones]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR38667.1 | 3.27e-135 | 3 | 541 | 6 | 554 |
| AIF49718.1 | 5.46e-53 | 22 | 346 | 18 | 325 |
| AIF51770.1 | 9.42e-52 | 28 | 346 | 23 | 324 |
| BBB92576.1 | 5.31e-51 | 29 | 343 | 25 | 324 |
| QCT94029.1 | 1.50e-50 | 6 | 331 | 3 | 317 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EZM_A | 1.86e-19 | 34 | 335 | 54 | 354 | CrystalStructure of ArnT from Cupriavidus metallidurans in the apo state [Cupriavidus metallidurans CH34],5F15_A Crystal Structure of ArnT from Cupriavidus metallidurans bound to Undecaprenyl phosphate [Cupriavidus metallidurans CH34] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O67270 | 2.31e-32 | 23 | 411 | 14 | 396 | Uncharacterized protein aq_1220 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1220 PE=3 SV=1 |
| O67601 | 7.90e-24 | 17 | 351 | 8 | 314 | Uncharacterized protein aq_1704 OS=Aquifex aeolicus (strain VF5) OX=224324 GN=aq_1704 PE=3 SV=1 |
| Q8D339 | 1.01e-22 | 2 | 332 | 9 | 330 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Wigglesworthia glossinidia brevipalpis OX=36870 GN=arnT PE=3 SV=1 |
| B4EUL1 | 1.28e-21 | 34 | 424 | 31 | 413 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 1 OS=Proteus mirabilis (strain HI4320) OX=529507 GN=arnT1 PE=3 SV=1 |
| B4ETL9 | 3.09e-21 | 17 | 332 | 14 | 323 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 2 OS=Proteus mirabilis (strain HI4320) OX=529507 GN=arnT2 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
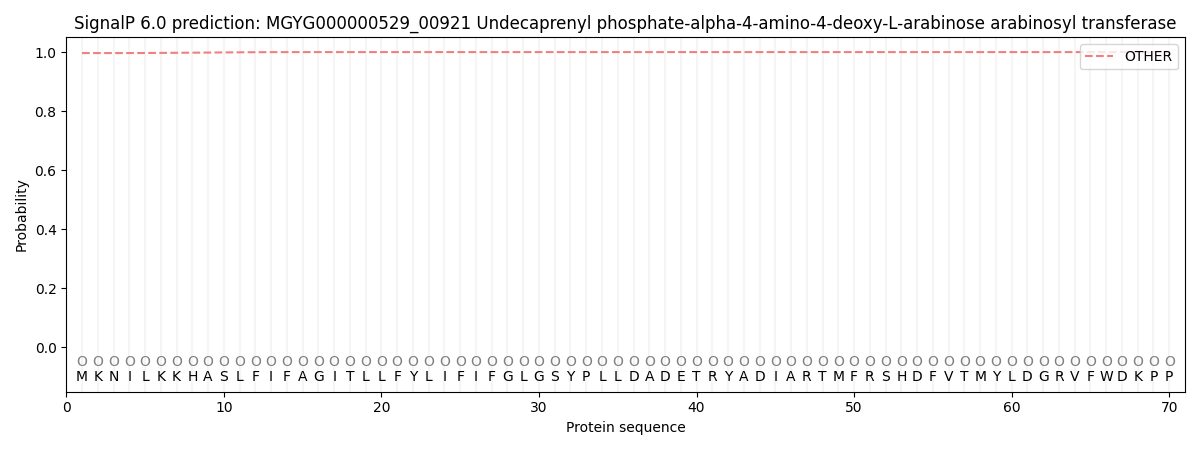
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.996698 | 0.000066 | 0.000024 | 0.000001 | 0.000000 | 0.003242 |
TMHMM Annotations download full data without filtering help
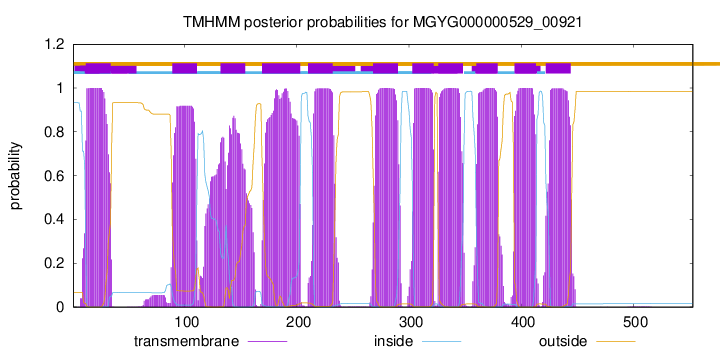
| start | end |
|---|---|
| 12 | 34 |
| 89 | 111 |
| 132 | 154 |
| 169 | 203 |
| 210 | 232 |
| 268 | 290 |
| 303 | 322 |
| 326 | 348 |
| 360 | 379 |
| 394 | 413 |
| 422 | 444 |
