You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000534_00092
You are here: Home > Sequence: MGYG000000534_00092
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900543975 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900543975 | |||||||||||
| CAZyme ID | MGYG000000534_00092 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 127857; End: 130100 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 152 | 407 | 1.8e-42 | 0.9583333333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00933 | Glyco_hydro_3 | 4.36e-20 | 129 | 406 | 40 | 280 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK15098 | PRK15098 | 3.77e-13 | 188 | 498 | 124 | 408 | beta-glucosidase BglX. |
| pfam01915 | Glyco_hydro_3_C | 2.09e-09 | 485 | 734 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QEC66867.1 | 1.35e-295 | 13 | 747 | 34 | 772 |
| AYL99540.1 | 5.07e-295 | 3 | 746 | 17 | 765 |
| QHS54798.1 | 1.06e-290 | 6 | 746 | 24 | 770 |
| QEM13852.1 | 6.51e-290 | 3 | 746 | 24 | 772 |
| AFS34656.1 | 6.98e-290 | 3 | 746 | 26 | 774 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5Z87_A | 3.17e-22 | 53 | 500 | 36 | 432 | ChainA, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
| 5JP0_A | 1.15e-20 | 188 | 735 | 127 | 638 | Bacteroidesovatus Xyloglucan PUL GH3B with bound glucose [Bacteroides ovatus],5JP0_B Bacteroides ovatus Xyloglucan PUL GH3B with bound glucose [Bacteroides ovatus] |
| 7EAP_A | 3.81e-15 | 53 | 498 | 7 | 402 | ChainA, Fn3_like domain-containing protein [Aspergillus oryzae RIB40] |
| 5YOT_A | 3.81e-15 | 53 | 498 | 7 | 402 | Isoprimeverose-producingenzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YOT_B Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YQS_A Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40],5YQS_B Isoprimeverose-producing enzyme from Aspergillus oryzae in complex with isoprimeverose [Aspergillus oryzae RIB40] |
| 6JGL_A | 1.62e-14 | 188 | 596 | 117 | 496 | Crystalstructure of barley exohydrolaseI W434H mutant in complex with methyl 2-thio-beta-sophoroside [Hordeum vulgare subsp. vulgare],6JGN_A Crystal structure of barley exohydrolaseI W434H in complex with 4'-nitrophenyl thiolaminaribioside [Hordeum vulgare subsp. vulgare],6JGO_A Crystal structure of barley exohydrolaseI W434H mutant in complex with 4I,4III,4V-S-trithiocellohexaose [Hordeum vulgare subsp. vulgare],6JGP_A Crystal structure of barley exohydrolaseI W434H mutant in complex with methyl 6-thio-beta-gentiobioside. [Hordeum vulgare subsp. vulgare] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q46684 | 4.70e-85 | 30 | 732 | 35 | 652 | Periplasmic beta-glucosidase/beta-xylosidase OS=Dickeya chrysanthemi OX=556 GN=bgxA PE=3 SV=1 |
| Q2UFP8 | 2.37e-84 | 53 | 735 | 46 | 627 | Probable beta-glucosidase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bglC PE=3 SV=2 |
| B8NGU6 | 5.99e-83 | 53 | 735 | 42 | 623 | Probable beta-glucosidase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglC PE=3 SV=1 |
| Q5BCC6 | 1.14e-81 | 53 | 734 | 38 | 615 | Beta-glucosidase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglC PE=1 SV=1 |
| T2KMH9 | 6.28e-24 | 53 | 735 | 29 | 639 | Putative beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22230 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
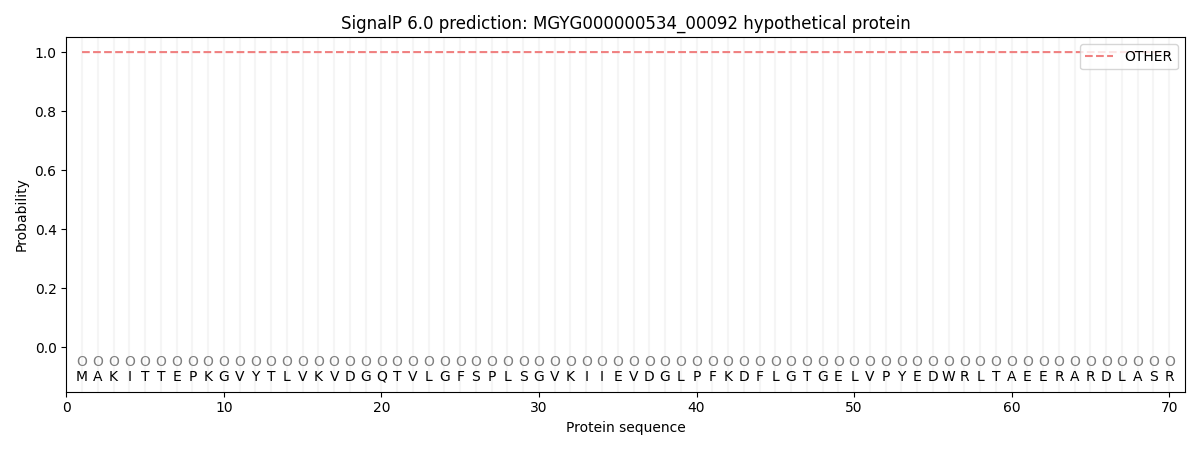
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000046 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
