You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000536_01996
You are here: Home > Sequence: MGYG000000536_01996
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cetobacterium_A sp900766645 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Fusobacteriota; Fusobacteriia; Fusobacteriales; Fusobacteriaceae; Cetobacterium_A; Cetobacterium_A sp900766645 | |||||||||||
| CAZyme ID | MGYG000000536_01996 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 497; End: 1258 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH73 | 134 | 248 | 2.6e-20 | 0.8828125 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2992 | Bax | 1.30e-30 | 66 | 251 | 62 | 259 | Uncharacterized FlgJ-related protein [General function prediction only]. |
| PRK10356 | PRK10356 | 4.61e-15 | 9 | 246 | 16 | 266 | protein bax. |
| pfam01832 | Glucosaminidase | 7.43e-12 | 137 | 195 | 17 | 77 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
| COG1705 | FlgJ | 1.29e-09 | 134 | 251 | 64 | 187 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| smart00047 | LYZ2 | 5.95e-08 | 134 | 211 | 29 | 114 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVQ19451.1 | 1.55e-62 | 22 | 251 | 49 | 284 |
| BBA51488.1 | 5.16e-58 | 26 | 251 | 31 | 263 |
| AVQ32154.1 | 1.46e-57 | 26 | 251 | 31 | 263 |
| AVQ29283.1 | 6.79e-56 | 26 | 251 | 32 | 264 |
| SQJ02627.1 | 6.79e-56 | 26 | 251 | 32 | 264 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P27297 | 9.21e-09 | 66 | 211 | 76 | 225 | Protein bax OS=Escherichia coli (strain K12) OX=83333 GN=bax PE=4 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as SP
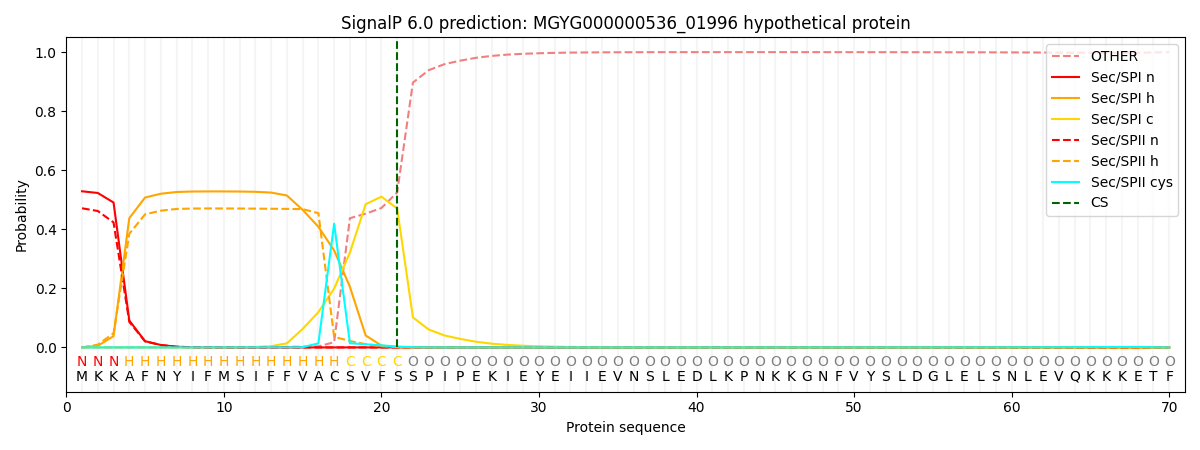
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001172 | 0.518681 | 0.479369 | 0.000227 | 0.000276 | 0.000253 |
