You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000538_01075
You are here: Home > Sequence: MGYG000000538_01075
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA1829 sp900549045 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; UBA1829; UBA1829 sp900549045 | |||||||||||
| CAZyme ID | MGYG000000538_01075 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 217990; End: 219723 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 10 | 533 | 1.2e-103 | 0.9203703703703704 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK13279 | arnT | 4.51e-111 | 10 | 537 | 8 | 508 | lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
| COG1807 | ArnT | 7.23e-46 | 1 | 543 | 1 | 524 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| pfam02366 | PMT | 4.78e-14 | 12 | 241 | 7 | 245 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This is a family of Dolichyl-phosphate-mannose-protein mannosyltransferase proteins EC:2.4.1.109. These proteins are responsible for O-linked glycosylation of proteins, they catalyze the reaction:- Dolichyl phosphate D-mannose + protein <=> dolichyl phosphate + O-D-mannosyl-protein. Also in this family is Drosophila rotated abdomen protein which is a putative mannosyltransferase. This family appears to be distantly related to pfam02516 (A Bateman pers. obs.). This family also contains sequences from ArnTs (4-amino-4-deoxy-L-arabinose lipid A transferase). They catalyze the addition of 4-amino-4-deoxy-l-arabinose (l-Ara4N) to the lipid A moiety of the lipopolysaccharide. This is a critical modification enabling bacteria (e.g. Escherichia coli and Salmonella typhimurium) to resist killing by antimicrobial peptides such as polymyxins. Members such as undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase are predicted to have 12 trans-membrane regions. The N-terminal portion of these proteins is hypothesized to have a conserved glycosylation activity which is shared between distantly related oligosaccharyltransferases ArnT and PglB families. |
| pfam13231 | PMT_2 | 1.18e-11 | 64 | 209 | 2 | 139 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
| COG1287 | Stt3 | 3.20e-05 | 65 | 211 | 96 | 244 | Asparagine N-glycosylation enzyme, membrane subunit Stt3 [Posttranslational modification, protein turnover, chaperones]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSH42095.1 | 2.41e-108 | 13 | 577 | 8 | 549 |
| AVM46372.1 | 2.12e-98 | 12 | 555 | 6 | 529 |
| QTF09713.1 | 5.41e-88 | 11 | 566 | 8 | 536 |
| VEA65444.1 | 2.22e-87 | 12 | 560 | 11 | 536 |
| QDW33282.1 | 4.37e-87 | 1 | 565 | 1 | 537 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EZM_A | 1.33e-38 | 18 | 466 | 42 | 466 | CrystalStructure of ArnT from Cupriavidus metallidurans in the apo state [Cupriavidus metallidurans CH34],5F15_A Crystal Structure of ArnT from Cupriavidus metallidurans bound to Undecaprenyl phosphate [Cupriavidus metallidurans CH34] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B4ETL9 | 1.38e-85 | 12 | 565 | 13 | 536 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase 2 OS=Proteus mirabilis (strain HI4320) OX=529507 GN=arnT2 PE=3 SV=1 |
| Q66A06 | 1.61e-83 | 1 | 565 | 1 | 537 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Yersinia pseudotuberculosis serotype I (strain IP32953) OX=273123 GN=arnT PE=2 SV=1 |
| A7FHH6 | 1.61e-83 | 1 | 565 | 1 | 537 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Yersinia pseudotuberculosis serotype O:1b (strain IP 31758) OX=349747 GN=arnT PE=3 SV=1 |
| Q1CIH9 | 1.61e-83 | 1 | 565 | 1 | 537 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Yersinia pestis bv. Antiqua (strain Nepal516) OX=377628 GN=arnT PE=3 SV=1 |
| A9R091 | 1.61e-83 | 1 | 565 | 1 | 537 | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase OS=Yersinia pestis bv. Antiqua (strain Angola) OX=349746 GN=arnT PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
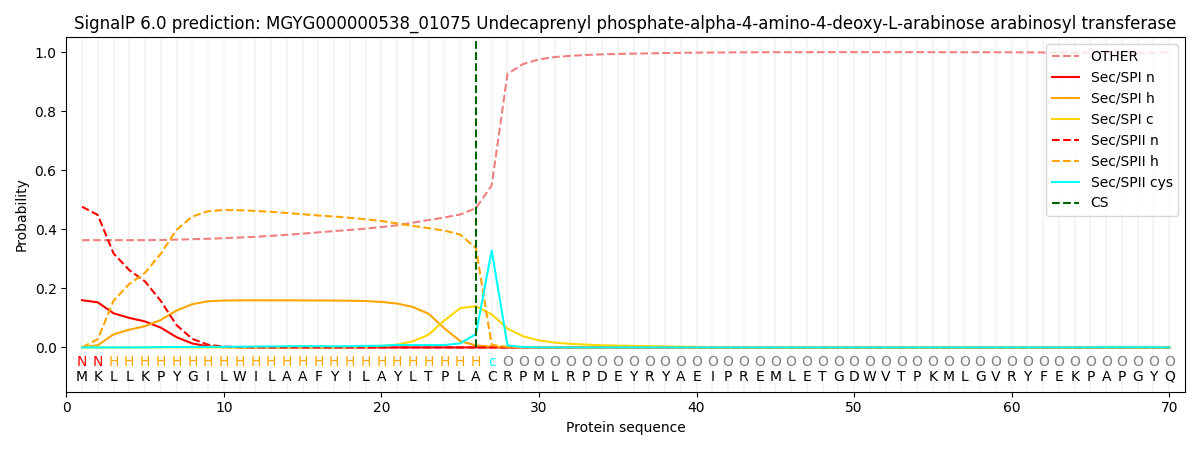
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.369422 | 0.149043 | 0.479651 | 0.000410 | 0.000370 | 0.001106 |
TMHMM Annotations download full data without filtering help
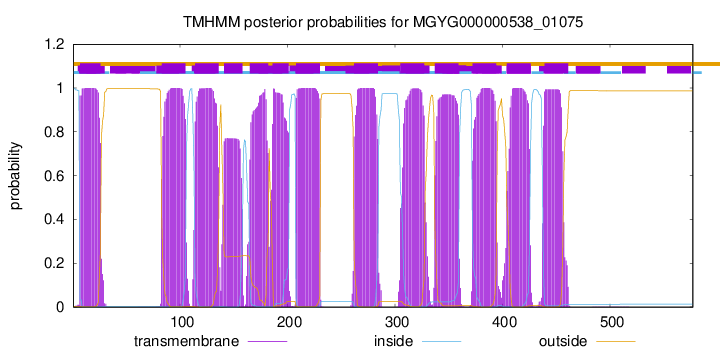
| start | end |
|---|---|
| 7 | 26 |
| 83 | 105 |
| 114 | 136 |
| 141 | 158 |
| 165 | 182 |
| 186 | 201 |
| 208 | 230 |
| 262 | 284 |
| 305 | 327 |
| 337 | 359 |
| 372 | 394 |
| 404 | 426 |
| 439 | 461 |
