You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000543_01442
You are here: Home > Sequence: MGYG000000543_01442
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Agathobacter sp900552085 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Agathobacter; Agathobacter sp900552085 | |||||||||||
| CAZyme ID | MGYG000000543_01442 | |||||||||||
| CAZy Family | GT8 | |||||||||||
| CAZyme Description | Glycosyltransferase-stabilizing protein Gtf2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10089; End: 11435 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR02919 | TIGR02919 | 5.89e-170 | 11 | 439 | 2 | 426 | accessory Sec system glycosyltransferase GtfB. Members of this protein family are found only in Gram-positive bacteria of the Firmicutes lineage, including several species of Staphylococcus, Streptococcus, and Lactobacillus. [Protein fate, Protein modification and repair] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK91596.1 | 1.11e-162 | 7 | 441 | 11 | 445 |
| CBK92843.1 | 2.23e-162 | 7 | 441 | 11 | 445 |
| ACR76237.1 | 4.49e-162 | 7 | 441 | 11 | 445 |
| QFJ55229.1 | 1.76e-151 | 8 | 435 | 7 | 432 |
| SNV42284.1 | 4.48e-127 | 10 | 442 | 1 | 430 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5E9U_B | 8.47e-111 | 11 | 435 | 2 | 425 | Crystalstructure of GtfA/B complex bound to UDP and GlcNAc [Streptococcus gordonii],5E9U_D Crystal structure of GtfA/B complex bound to UDP and GlcNAc [Streptococcus gordonii],5E9U_F Crystal structure of GtfA/B complex bound to UDP and GlcNAc [Streptococcus gordonii],5E9U_H Crystal structure of GtfA/B complex bound to UDP and GlcNAc [Streptococcus gordonii] |
| 5E9T_B | 8.73e-111 | 11 | 435 | 2 | 425 | Crystalstructure of GtfA/B complex [Streptococcus gordonii],5E9T_D Crystal structure of GtfA/B complex [Streptococcus gordonii] |
| 5GVW_A | 1.23e-10 | 286 | 406 | 269 | 389 | Crystalstructure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_B Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_C Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVW_D Crystal structure of the apo-form glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae] |
| 5GVV_A | 9.07e-10 | 286 | 406 | 269 | 389 | Crystalstructure of the glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae],5GVV_F Crystal structure of the glycosyltransferase GlyE in Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q3S2Y1 | 1.14e-123 | 10 | 422 | 1 | 410 | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase stabilizing protein GtfB OS=Streptococcus agalactiae OX=1311 GN=gtfB PE=1 SV=1 |
| Q79T00 | 4.55e-114 | 10 | 435 | 1 | 425 | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase stabilizing protein GtfB OS=Streptococcus gordonii OX=1302 GN=gtfB PE=1 SV=1 |
| A1C3M0 | 1.07e-106 | 10 | 407 | 1 | 394 | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase stabilizing protein GtfB OS=Streptococcus parasanguinis OX=1318 GN=gtfB PE=1 SV=1 |
| A0A0H2UR90 | 3.64e-106 | 10 | 439 | 1 | 427 | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase stabilizing protein GtfB OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=gtfB PE=1 SV=1 |
| A0A0S4NND9 | 2.15e-74 | 13 | 439 | 4 | 423 | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase stabilizing protein GtfB OS=Limosilactobacillus reuteri (strain ATCC 53608) OX=927703 GN=gtfB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
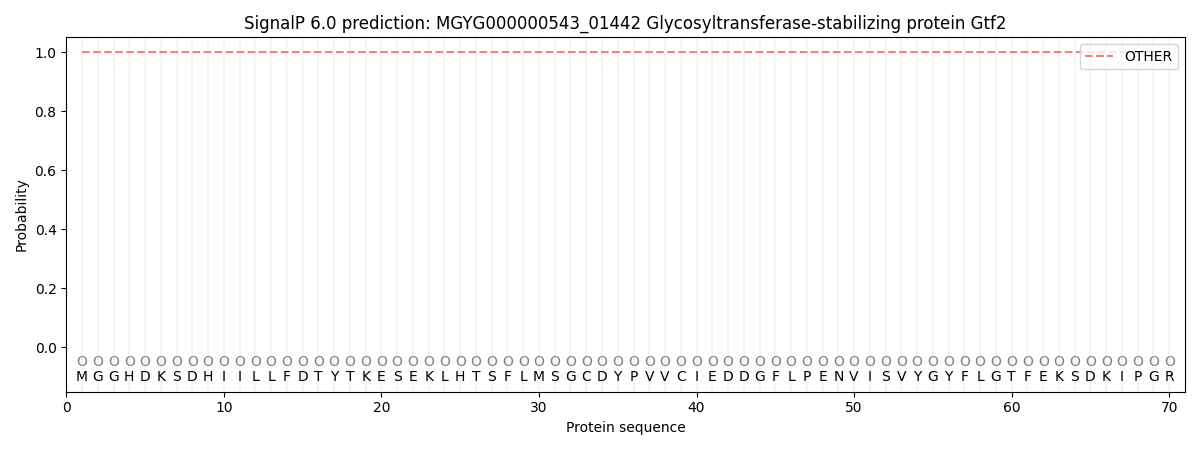
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000062 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
