You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000544_00341
You are here: Home > Sequence: MGYG000000544_00341
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Zag1 sp900769355 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Cyanobacteria; Vampirovibrionia; Gastranaerophilales; Gastranaerophilaceae; Zag1; Zag1 sp900769355 | |||||||||||
| CAZyme ID | MGYG000000544_00341 | |||||||||||
| CAZy Family | CE11 | |||||||||||
| CAZyme Description | UDP-3-O-acyl-N-acetylglucosamine deacetylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14023; End: 14805 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE11 | 2 | 252 | 6.7e-61 | 0.9298892988929889 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03331 | LpxC | 6.85e-71 | 4 | 252 | 19 | 269 | UDP-3-O-acyl N-acetylglycosamine deacetylase. The enzymes in this family catalyze the second step in the biosynthetic pathway for lipid A. |
| PRK13186 | lpxC | 8.10e-67 | 2 | 252 | 20 | 272 | UDP-3-O-acyl-N-acetylglucosamine deacetylase. |
| COG0774 | LpxC | 1.13e-62 | 4 | 252 | 22 | 275 | UDP-3-O-acyl-N-acetylglucosamine deacetylase [Cell wall/membrane/envelope biogenesis]. |
| PRK13188 | PRK13188 | 4.45e-45 | 4 | 260 | 23 | 306 | bifunctional UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase/(3R)-hydroxymyristoyl-[acyl-carrier-protein] dehydratase; Reviewed |
| TIGR00325 | lpxC | 1.03e-43 | 2 | 250 | 19 | 269 | UDP-3-0-acyl N-acetylglucosamine deacetylase. UDP-3-O-(R-3-hydroxymyristoyl)-GlcNAc deacetylase from E. coli , LpxC, was previously designated EnvA. This enzyme is involved in lipid-A precursor biosynthesis. It is essential for cell viability. [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR38878.1 | 7.36e-181 | 1 | 260 | 51 | 310 |
| AGB42301.1 | 6.38e-38 | 30 | 257 | 53 | 275 |
| BBH38876.1 | 1.12e-37 | 8 | 254 | 24 | 270 |
| BAG03744.1 | 1.12e-37 | 8 | 254 | 24 | 270 |
| BCU10606.1 | 3.10e-37 | 8 | 254 | 24 | 270 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2GO3_A | 1.70e-24 | 12 | 252 | 30 | 263 | ChainA, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2GO3_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2GO4_A Chain A, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2GO4_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus] |
| 1P42_A | 1.80e-24 | 12 | 252 | 29 | 262 | ChainA, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],1P42_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],1YH8_A Chain A, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],1YH8_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],1YHC_A Chain A, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],1YHC_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus] |
| 2IER_A | 1.84e-24 | 12 | 252 | 30 | 263 | ChainA, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2IER_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2IES_A Chain A, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2IES_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2J65_A Structure of LpxC from Aquifex aeolicus in complex with UDP [Aquifex aeolicus VF5],2J65_B Structure of LpxC from Aquifex aeolicus in complex with UDP [Aquifex aeolicus VF5],2O3Z_A Chain A, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],2O3Z_B Chain B, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],3P76_A X-ray crystal structure of Aquifex aeolicus LpxC complexed SCH1379777 [Aquifex aeolicus],4U3B_A LpxC from A.Aaeolicus in complex with the MMP inhibitor 4-[[4-(4-chlorophenoxy)phenyl]sulfanylmethyl]tetrahydropyran-4-carbohydroxamic acid - compound 2 [Aquifex aeolicus VF5],4U3D_A LpxC from A.Aaeolicus in complex with 4-[[4-[2-[4-(morpholinomethyl)phenyl]ethynyl]phenoxy]methyl]tetrahydropyran-4-carbohydroxamic acid (compound 9) [Aquifex aeolicus VF5] |
| 2JT2_A | 1.94e-24 | 12 | 252 | 30 | 263 | ChainA, UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase [Aquifex aeolicus],5DRO_A Structure of the Aquifex aeolicus LpxC/LPC-011 Complex [Aquifex aeolicus VF5],5DRO_B Structure of the Aquifex aeolicus LpxC/LPC-011 Complex [Aquifex aeolicus VF5],5DRP_A Structure of the AaLpxC/LPC-023 Complex [Aquifex aeolicus VF5],5DRP_B Structure of the AaLpxC/LPC-023 Complex [Aquifex aeolicus VF5] |
| 3P3C_A | 1.94e-24 | 12 | 252 | 29 | 262 | CrystalStructure of the Aquifex aeolicus LpxC/LPC-009 complex [Aquifex aeolicus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B0SS82 | 2.02e-36 | 4 | 252 | 25 | 275 | UDP-3-O-acyl-N-acetylglucosamine deacetylase OS=Leptospira biflexa serovar Patoc (strain Patoc 1 / ATCC 23582 / Paris) OX=456481 GN=lpxC PE=3 SV=1 |
| B0S9V0 | 2.02e-36 | 4 | 252 | 25 | 275 | UDP-3-O-acyl-N-acetylglucosamine deacetylase OS=Leptospira biflexa serovar Patoc (strain Patoc 1 / Ames) OX=355278 GN=lpxC PE=3 SV=1 |
| B2IWK6 | 8.74e-36 | 8 | 252 | 25 | 277 | UDP-3-O-acyl-N-acetylglucosamine deacetylase OS=Nostoc punctiforme (strain ATCC 29133 / PCC 73102) OX=63737 GN=lpxC PE=3 SV=1 |
| B7KKQ2 | 8.95e-36 | 8 | 254 | 25 | 274 | UDP-3-O-acyl-N-acetylglucosamine deacetylase OS=Gloeothece citriformis (strain PCC 7424) OX=65393 GN=lpxC PE=3 SV=1 |
| Q8F3U4 | 5.82e-35 | 34 | 252 | 59 | 274 | UDP-3-O-acyl-N-acetylglucosamine deacetylase OS=Leptospira interrogans serogroup Icterohaemorrhagiae serovar Lai (strain 56601) OX=189518 GN=lpxC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
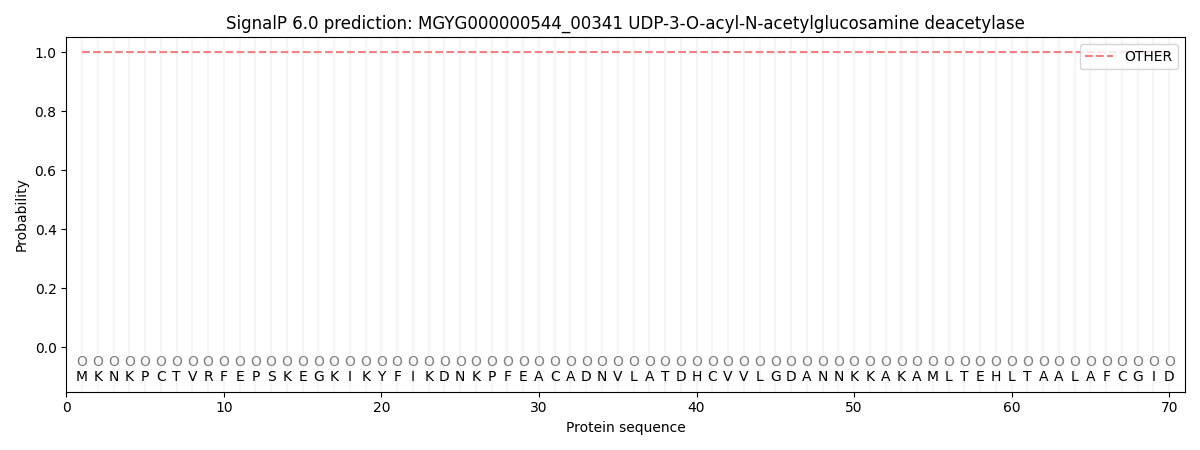
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
