You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000545_00793
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Mediterraneibacter;
|
| CAZyme ID |
MGYG000000545_00793
|
| CAZy Family |
GH140 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000000545 |
2021811 |
MAG |
Fiji |
Oceania |
|
| Gene Location |
Start: 14996;
End: 16252
Strand: -
|
No EC number prediction in MGYG000000545_00793.
| Family |
Start |
End |
Evalue |
family coverage |
| GH140 |
2 |
380 |
3.8e-106 |
0.912621359223301 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam13204
|
DUF4038 |
3.59e-105 |
13 |
328 |
8 |
320 |
Protein of unknown function (DUF4038). A family of putative cellulases. |
This protein is predicted as OTHER
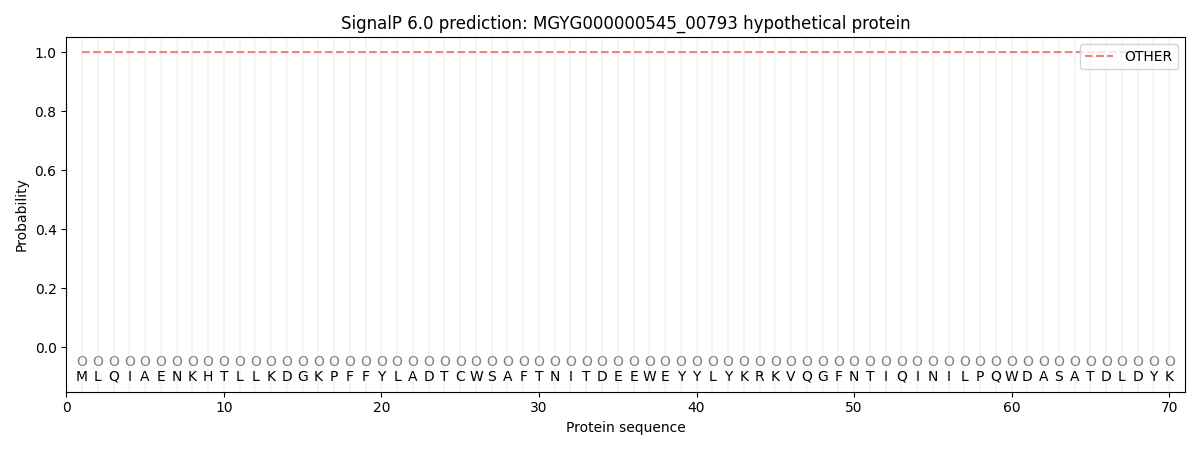
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000047
|
0.000001
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|