You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000553_00067
You are here: Home > Sequence: MGYG000000553_00067
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
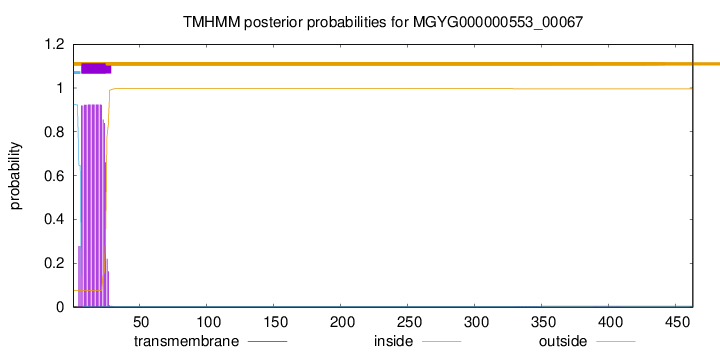
TMHMM annotations
Basic Information help
| Species | Prevotella sp900548535 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900548535 | |||||||||||
| CAZyme ID | MGYG000000553_00067 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 91479; End: 92870 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00118 | LysM | 6.74e-09 | 30 | 75 | 2 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| smart00257 | LysM | 6.83e-08 | 30 | 75 | 1 | 44 | Lysin motif. |
| pfam01476 | LysM | 2.42e-07 | 31 | 76 | 1 | 43 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
| COG1388 | LysM | 0.001 | 30 | 88 | 68 | 122 | LysM repeat [Cell wall/membrane/envelope biogenesis]. |
| cd14528 | PFA-DSP_Siw14 | 0.010 | 197 | 272 | 50 | 115 | atypical dual specificity phosphatases similar to yeast Siw14. This subfamily contains Saccharomyces Siw14 and a novel phosphatase from the Arabidopsis thaliana gene locus At1g05000. Siw14, also known as Oca3, plays a role in actin filament organization and endocytosis. Siw14 has been shown to be an inositol pyrophosphate phosphatase, hydrolyzing the beta-phosphate from 5-diphosphoinositol pentakisphosphate (5PP-IP5or IP7). The At1g05000 protein, also called AtPFA-DSP1, has been shown to have highest activity toward olyphosphate (poly-P(12-13)) and deoxyribo- and ribonucleoside triphosphates, and less activity toward phosphoenolpyruvate, phosphotyrosine, phosphotyrosine-containing peptides, and phosphatidylinositols. This subfamily belongs to a group of atypical DSPs present in plants, fungi, kinetoplastids, and slime molds called plant and fungi atypical dual-specificity phosphatases (PFA-DSPs). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB46898.1 | 1.20e-186 | 6 | 463 | 7 | 451 |
| ALO48876.1 | 1.73e-168 | 1 | 463 | 1 | 448 |
| QUB62527.1 | 5.84e-163 | 1 | 463 | 1 | 463 |
| QUB56650.1 | 8.28e-163 | 1 | 463 | 1 | 463 |
| QUB57976.1 | 8.28e-163 | 1 | 463 | 1 | 463 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.021903 | 0.546119 | 0.431024 | 0.000319 | 0.000286 | 0.000335 |